- 1. TG-263 and the common language of radiation therapy
- 2. What is TG-263
- 3. Real-world problems caused by lack of standardization
- 4. How to apply TG-263 in clinical practice
- 5. Complete TG-263 nomenclature table 717 structures
- 6. Integration with treatment planning systems and artificial intelligence
- 7. Implementation workflow in 10 steps
- 8. Relationship with target volume delineation
- 9. The future of standardization in radiation therapy
- 10. Conclusion
TG-263 and the common language of radiation therapy
A medical physicist receives a treatment plan exported from another institution. Upon opening the DICOM file, the left parotid is labeled Lt Parotid. In the next case, from the same center, the name appears as PAROTID_L. A third patient shows parotid_left. And in the receiving department’s own template, the same structure is listed as L_Parotid_Gland. Five different names for the same organ at risk, within a relatively small universe of cases.
This scenario is not hypothetical. It reproduces real data collected by the AAPM prior to the publication of TG-263. Across 20 surveyed institutions, the task group found between 21 and 311 defined structure names, with variations ranging from personal abbreviations to entire conventions created by a single professional and never formally documented.
The problem seems minor when viewed in isolation. But it accumulates fast. When a department tries to automate DVH review, train an auto-segmentation model, consolidate data for a multicenter study, or simply compare plans across patients treated in different years, the lack of standardization becomes constant rework. Every analysis requires a preliminary “translation” step that consumes time, introduces errors, and prevents real automation.
The AAPM recognized this bottleneck and, in 2012, established Task Group 263 with a clear mandate: create a nomenclature standard for anatomical structures, target volumes, and DVH metrics that could be adopted by any radiation therapy department, regardless of vendor, country, or operational scale. The group brought together 57 participants — 33 clinical physicists, 15 radiation oncologists, 8 vendor representatives, and 1 dosimetrist — and worked for five years until the report was published in January 2018.
The result is more than a list of names. TG-263 defines nomenclature construction principles, laterality rules, conventions for target volumes and their dose levels, standardized notation for dosimetric metrics, and a 10-step implementation workflow that allows any department to adopt the standard without interrupting clinical operations.
This guide covers the full content of the report, includes the complete table with all 717 cataloged non-target structures, explains how the principles apply in daily practice, and details the integration with treatment planning systems, artificial intelligence tools, and multicenter research workflows. The goal is to serve as a comprehensive reference for any professional who needs to implement, audit, or expand TG-263 nomenclature in their department.
What is TG-263
The AAPM (American Association of Physicists in Medicine) Task Group 263 was born from the realization that radiation therapy needed a shared technical vocabulary for naming anatomical structures in treatment planning systems. Before this standard, each institution developed its own conventions — when it developed any at all. The result was a fragmentation that hindered any activity dependent on consistent data across patients, equipment, or centers.
The group was formally constituted in 2012 with members nominated by both the AAPM and ASTRO (American Society for Radiation Oncology). The final composition included 39 AAPM members and 41 ASTRO members, with overlap between the two societies, totaling 57 unique participants. The diversity of the group was intentional: the presence of physicists, physicians, dosimetrists, and vendor representatives ensured that the standard would be viable from both clinical and technological standpoints.
The report was published in January 2018 as AAPM Report No. 263, accompanied by a peer-reviewed publication in Medical Physics (the 2018 S0360 article). The main document defines the principles; the supplemental material, available on the AAPM website, contains the complete spreadsheet with all cataloged structures.
Spreadsheet structure
The core of TG-263 is a spreadsheet organized into 9 columns that classify each structure systematically. This organization is not arbitrary — each column serves a specific purpose in the clinical and informatic use chain:
Target Type distinguishes whether the structure is anatomic, a target volume (GTV, CTV, PTV), a PRV, or a derived planning structure. Major Category groups by broad anatomic system — bone, nerve, organ, vessel, lymph node. Minor Category refines this grouping with more specific subcategories. Anatomic Group indicates the body region: head and neck, thorax, abdomen, pelvis, body, limb.
N Characters records the length of the standardized name, a critical piece of information because TG-263 recommends a maximum of 16 characters to ensure compatibility with legacy systems and interface fields. TG263-Primary Name is the principal name to be used in the treatment planning system — for example, Parotid_L for the left parotid. TG-263-Reverse Order Name provides a reversed version (L_Parotid) to facilitate search and sorting in long lists.
Description provides the full anatomical name without abbreviations. FMAID maps the structure to the Foundational Model of Anatomy, an anatomical ontology maintained by the University of Washington that enables interoperability with SNOMED CT, DICOM, and other health informatics standards.
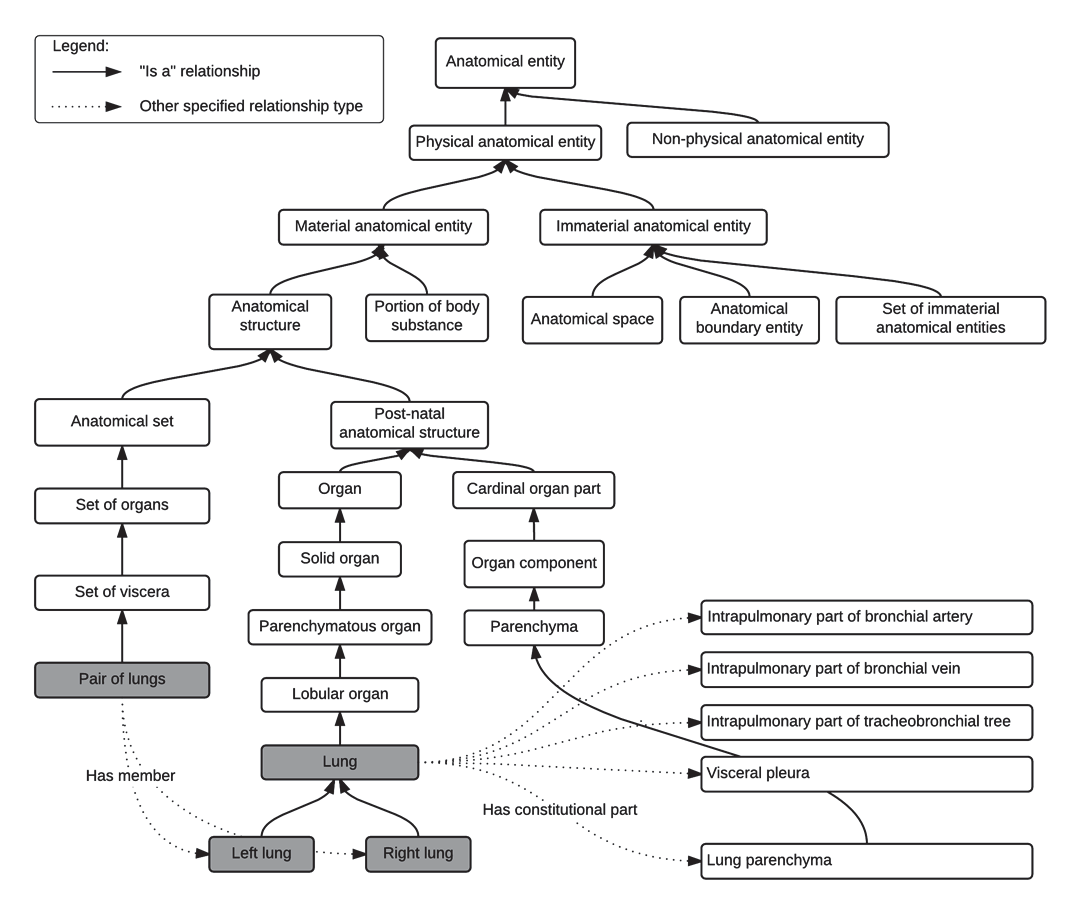
Primary Name versus Reverse Order Name
The distinction between these two fields is one of the most important concepts in TG-263 and is frequently misunderstood. The Primary Name follows the Structure_Laterality pattern — for example, Kidney_L, Femur_R, CN_VII_L. This format facilitates human reading and follows the natural convention of naming the structure first and then qualifying it.
The Reverse Order Name inverts to Laterality_Structure — L_Kidney, R_Femur, L_CN_VII. This format is useful when the structure list needs to be sorted alphabetically by laterality, grouping all left-sided and all right-sided structures together. Some treatment planning systems and analysis tools benefit from this ordering for rapid review.
TG-263 recommends that the Primary Name be used as the preferred name in the TPS. The Reverse Order Name exists as a documented alternative, not a substitute. Departments adopting the standard should choose one convention and apply it consistently across all operations.
Scope: 717 non-target structures
The spreadsheet catalogs 717 non-target structures organized into 7 anatomic groups: Head and Neck (282), Thorax (180), Abdomen (60), Pelvis (109), Body/Systemic (52), Limb singular (16), and Limbs bilateral (17). This catalog covers virtually every anatomical structure that a radiation therapy department encounters in clinical practice, from cranial bones and nerves to vessels, lymph nodes, abdominal organs, and pelvic structures.
Beyond anatomical structures, TG-263 defines separate conventions for target volumes (GTV, CTV, ITV, PTV, and their variants), planning structures (PRV, auxiliary structures with the z prefix), and dosimetric metrics (DVH). These three domains follow their own principles, detailed in subsequent sections of this guide.
Real-world problems caused by lack of standardization
TG-263 did not arise from a theoretical concern. Before defining any standard, the group conducted a survey across 20 North American institutions to document the actual state of nomenclature in radiation therapy. The findings were revealing.
Of the 20 surveyed institutions, 16 reported having some form of defined nomenclature for at least some disease sites. But “having nomenclature” did not mean having consistency. The number of defined structures ranged from 21 to 311 across institutions, and within each one there were variations between professionals, between rooms, and across time periods. None of the 20 covered all anatomical structures needed for all treatment modalities.
The most cited example in the report is the left optic nerve. When collecting the names used across surveyed institutions, the group found 12 variants for a single structure: Lt Optic Nerve, L_Optic_Nerve, OpticNrv_L, Optic_Nerve_Left, among others. Each variant was internally logical to its creator, but incompatible with the rest when data needed to be combined.
Impact on data mining
A department that accumulates 10 years of treatment plans holds a valuable repository for research, benchmarking, and quality improvement. But if those plans use inconsistent names for the same structures, any database query becomes an exercise in manual mapping. Extracting “all mean doses to the spinal cord” requires first identifying whether the cord was named SpinalCord, Spinal_Cord, SC, MedulaEspinhal, Cord, or any other local variation.
This mapping cost is not trivial. In multicenter studies with thousands of patients, the nomenclature harmonization step can consume weeks of work and still contain errors. TG-263 eliminates this problem at its source: if all centers use SpinalCord, the database query returns complete results without manual intervention.
Impact on auto-segmentation and AI
Deep learning models for auto-segmentation require training data labeled consistently. A model trained on contours labeled Parotid_L cannot automatically use data labeled Lt_Parotid_Gland — the mapping must be done manually for each variant, and any error in that mapping contaminates the training.
When a department adopts TG-263 nomenclature, its data becomes immediately compatible with models trained at other departments following the same standard. This accelerates the adoption of auto-segmentation, improves training quality by enabling larger datasets, and makes cross-institutional validation feasible.
Impact on QA automation
Automated quality assurance protocols depend on predictable names. A script that verifies whether the maximum dose to the lens does not exceed the institutional limit needs to know exactly how the lens is named. If the name varies between Lens_L, L_Lens, Left_Lens, and Cristalino_E, the script needs a synonym table that grows with every new variant encountered — or, worse, fails silently when it encounters an uncataloged name.
TG-263 allows QA tools to be written once and work in any compliant department. The predictability of names is the foundation for reliable automation.
Impact on multicenter studies and clinical trials
Radiation therapy clinical trials that collect dosimetric data from multiple institutions face the nomenclature problem at scale. The RTOG (now NRG Oncology) published naming guidelines for specific studies, but without a universal standard adopted by the community, each new trial repeated the same cycle of definition, communication, and verification. TG-263 provides a permanent foundation that reduces this overhead for any future trial.
How to apply TG-263 in clinical practice
TG-263 defines two sets of principles: 15 for non-target structures (organs at risk and anatomical structures) and 10 for target volumes. It also standardizes dosimetric metric notation. Each set was designed to solve specific problems in clinical practice.
Principles for non-target structures
16-character limit. The standardized name must not exceed 16 characters. This limit is not arbitrary — it reflects real interface field constraints in both legacy and contemporary treatment planning systems. Longer names are truncated by some TPS, generating ambiguity. The limit enforces conciseness and prevents the proliferation of long descriptive names that no one types identically twice.
Case-insensitive uniqueness. Each name must be unique regardless of uppercase and lowercase. Brainstem and brainstem cannot coexist as distinct structures. This rule prevents errors in systems that ignore case during search or sorting.
No spaces, underscore as separator. Spaces are prohibited in TG-263 names. The standard separator is the underscore (_). Thus, Optic_Nerve_L and not Optic Nerve L. This convention avoids problems with parsers, scripts, and interfaces that treat spaces as field delimiters.
Laterality suffixes: _L, _R, _A, _P. Lateralized structures receive the corresponding suffix at the end of the name. The standard uses single letters for left (_L), right (_R), anterior (_A), and posterior (_P). Bilateral structures without defined laterality use the plural form — for example, Kidneys for both kidneys, Kidney_L and Kidney_R for each individually.
Category root prefixes. Structures from specific categories use standardized prefixes that facilitate grouping and search:
A_ for arteries (A_Carotid_L), V_ for veins (V_Jugular_R), LN_ for lymph nodes (LN_Ax_L1_L), CN_ for cranial nerves (CN_VII_L), Bone_ for bones (Bone_Mandible), Musc_ for muscles (Musc_Masseter_L). These prefixes ensure that all arteries are grouped together in the list, all lymph nodes together, and so on.
PRV structures. Planning Risk Volumes (PRV) follow the convention Structure_PRV for a generic margin or Structure_PRVxx where xx indicates the margin in millimeters. Examples: Brainstem_PRV for a brainstem PRV with an unspecified margin, SpinalCord_PRV05 for a spinal cord PRV with a 5 mm expansion. This convention allows any professional to immediately identify the base structure and the applied margin without consulting external documentation.
Partial structures: the tilde (~). When only part of a structure is delineated, TG-263 uses the tilde as a suffix: Brain~ for partial brain, Lung~_L for partial left lung. This situation is common in treatments where the acquisition or treatment field does not cover the entire structure — the tilde explicitly signals that coverage is incomplete, preventing DVH metrics from being interpreted as representative of the whole organ.
Planning structures: the z prefix. Structures created solely for plan optimization — rings, subtraction structures, dose auxiliaries — receive the z prefix so they appear at the end of the alphabetical list and do not mix with real anatomical structures. Examples: zPTVopt, zRing, zBolus. The convention is simple and effective: when a physicist scrolls through the structure list, everything starting with z is a planning auxiliary and can be filtered or ignored depending on the context.
Principles for target volumes
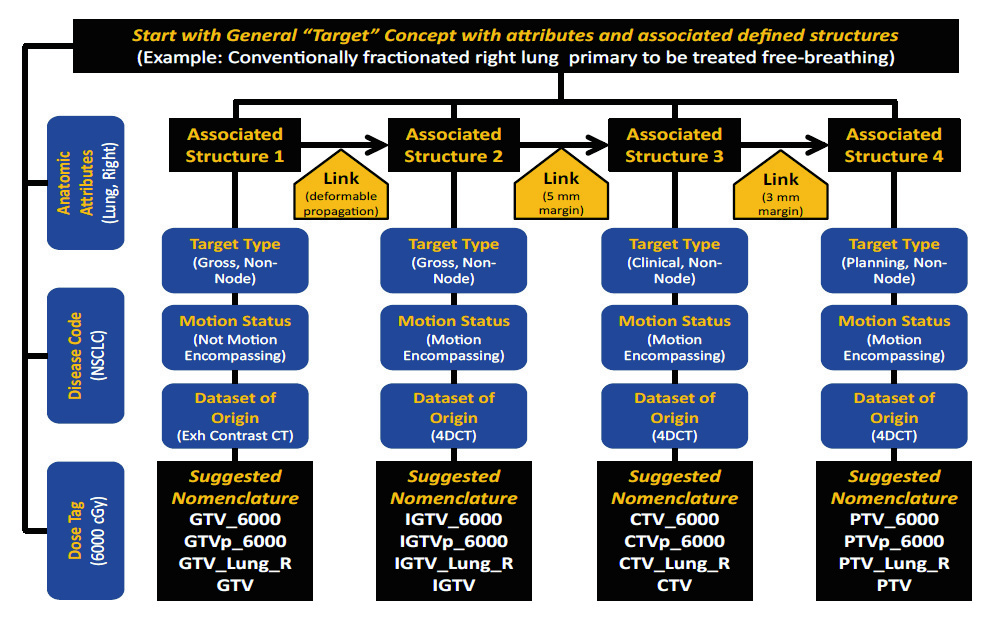
Target volume nomenclature follows 10 principles constructed to reflect the clinical hierarchy between gross tumor volume, clinical target volume, internal margins, and planning target volume.
Volume types. TG-263 defines the following types: GTV (Gross Tumor Volume), CTV (Clinical Target Volume), ITV (Internal Target Volume), IGTV (Internal Gross Target Volume), ICTV (Internal Clinical Target Volume), PTV (Planning Target Volume), and PTV! (evaluation PTV, decoupled from optimization constraints). Each type occupies a precise position in the clinical derivation chain.
Classifiers. Volumes may receive classifiers that indicate the nature of the disease: n for nodal (GTV_n), p for primary (CTV_p), sb for surgical bed (CTV_sb), par for parenchymal. These classifiers combine with the volume type and allow identification of the clinical origin without ambiguity. A plan with GTV_p, GTV_n, CTV_p, and CTV_n communicates in four names the entire coverage strategy for the primary tumor and nodal disease.
Dose levels. When the plan prescribes different doses to distinct volumes, TG-263 allows the level to be indicated in the name: PTV_High, PTV_Low, PTV_Mid for relative levels, or PTV_5040 for absolute dose in cGy. This convention is particularly useful in SIB (Simultaneous Integrated Boost) plans, where three or more dose levels coexist and clarity about which PTV receives which prescription is critical for treatment safety.
Target name composition. The elements combine in a predictable pattern: Type_Classifier_DoseLevel. Thus, PTV_n_High unambiguously identifies the nodal planning target volume receiving the highest dose. CTV_sb_Low is the clinical target volume of the surgical bed at the low dose level. The composition is modular and extensible without violating the construction rules.
Dosimetric metric nomenclature (DVH)
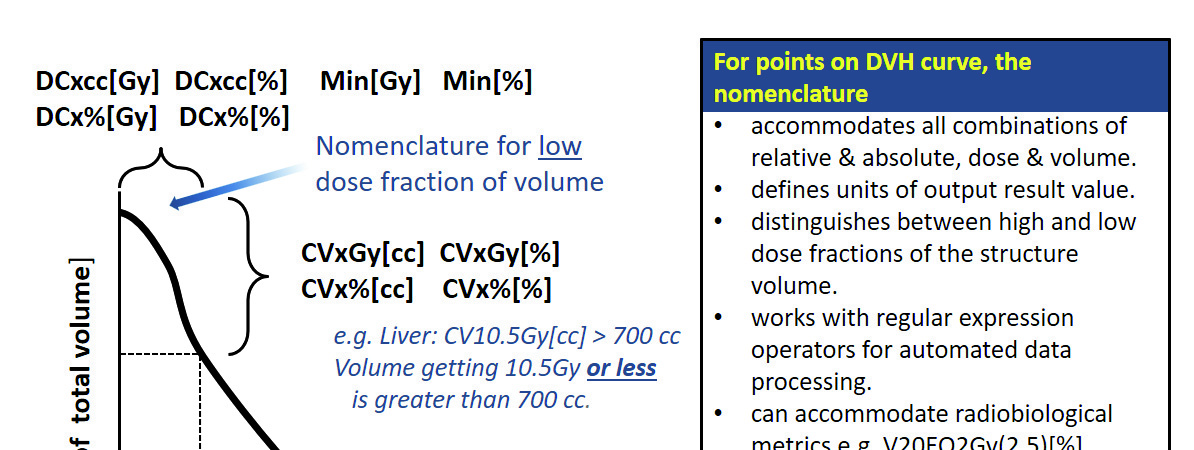

TG-263 standardizes how DVH metrics are written to eliminate ambiguity. The notation follows the format MetricValueUnit[ResultUnit]:
V20Gy[%] — volume receiving at least 20 Gy, expressed as a percentage. V20Gy[cc] — the same volume, expressed in cubic centimeters. D0.1cc[Gy] — dose covering at least 0.1 cc of the volume, in Gy. D95%[Gy] — dose at the point covering 95% of the volume. Mean[Gy] — mean dose to the volume. Max[Gy] — point maximum dose.
This notation resolves a chronic problem: the same metric written in different ways (V20, V_20Gy, V20_pct, Vol_20Gy_%) across reports, protocols, and scripts. With the TG-263 standard, a script that calculates V20Gy[%] produces output that is readable by any other script or professional familiar with the convention.
Complete TG-263 nomenclature table
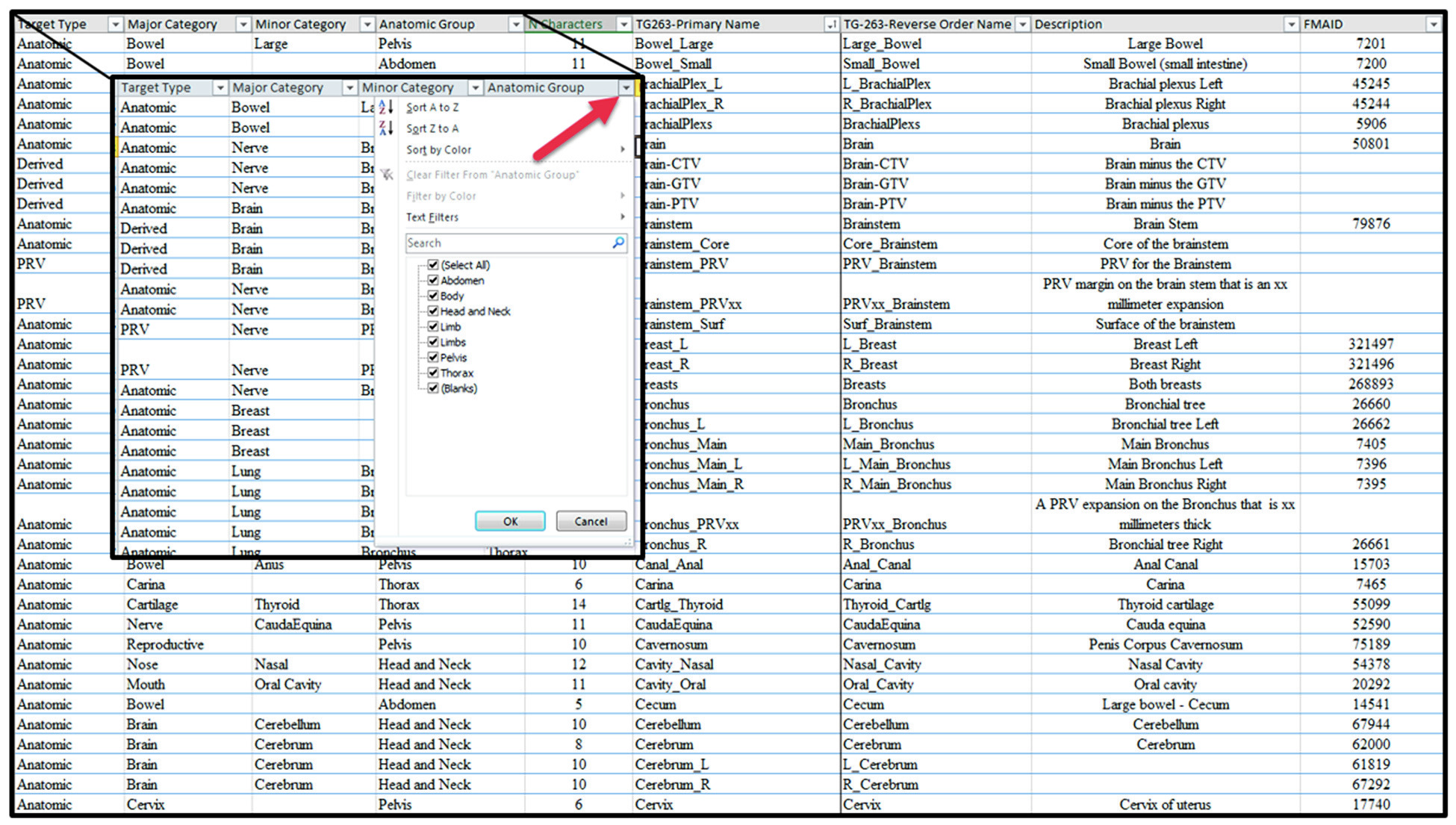
The table below compiles all 717 non-target structures cataloged by TG-263, organized into 7 anatomic groups. Use the search field to quickly locate any structure by name, category, or region. Names follow the TG263-Primary Name format from the official spreadsheet (AAPM TG-263 supplemental materials); the FMAID column indicates the Foundational Model of Anatomy identifier when available.
Head and Neck — 282 structures
| TG-263 Name | Type | Category | Description | FMAID |
|---|---|---|---|---|
| A_Carotid | Anatomic | Artery | Common Carotid Artery | 3939 |
| A_Carotid_L | Anatomic | Artery | Carotid Artery | 4058 |
| A_Carotid_R | Anatomic | Artery | Carotid Artery | 3941 |
| A_Coronary | Anatomic | Artery | Coronary Artery | 49893 |
| A_Hypophyseal_I | Anatomic | Artery | Hypophyseal Artery Inferior | 49846 |
| A_Hypophyseal_S | Anatomic | Artery | Hypophyseal Artery Superior | 49849 |
| Arytenoid | Anatomic | Cartilage | Arytenoid cartilage | 55109 |
| Arytenoid_L | Anatomic | Cartilage | Arytenoid cartilage Left | 55114 |
| Arytenoid_R | Anatomic | Cartilage | Arytenoid cartilage Right | 55113 |
| Bone_Ethmoid | Anatomic | Bone | Ethmoid Bone | 52740 |
| Bone_Frontal | Anatomic | Bone | Frontal Bone | 52734 |
| Bone_Hyoid | Anatomic | Bone | Hyoid Bone | 52749 |
| Bone_Incus | Anatomic | Ear | Incus | 52752 |
| Bone_Incus_L | Anatomic | Ear | Incus Left | 74051 |
| Bone_Incus_R | Anatomic | Ear | Incus Right | 74050 |
| Bone_Lacrimal | Anatomic | Bone | Lacrimal Bone | 52741 |
| Bone_Lacrimal_L | Anatomic | Bone | Lacrimal Bone Left | 53646 |
| Bone_Lacrimal_R | Anatomic | Bone | Lacrimal Bone Right | 53645 |
| Bone_Mandible | Anatomic | Bone | Mandible | 52748 |
| Bone_Mastoid | Anatomic | Bone | Both Mastoids | 52872 |
| Bone_Mastoid_L | Anatomic | Bone | Left Mastoid Bone | 52874 |
| Bone_Mastoid_R | Anatomic | Bone | Right Mastoid Bone | 52873 |
| Bone_Nasal | Anatomic | Bone | Nasal Bone | 52745 |
| Bone_Nasal_L | Anatomic | Bone | Nasal Bone Left | 53648 |
| Bone_Nasal_R | Anatomic | Bone | Nasal Bone Right | 53647 |
| Bone_Occipital | Anatomic | Bone | Occipital Bone | 52735 |
| Bone_Palatine | Anatomic | Bone | Palatine bone | 52746 |
| Bone_Palatine_L | Anatomic | Bone | Palatine bone Left | 53656 |
| Bone_Palatine_R | Anatomic | Bone | Palatine bone Right | 53655 |
| Bone_Parietal | Anatomic | Bone | Parietal bone | 9613 |
| Bone_Parietal_L | Anatomic | Bone | Parietal bone Left | 52789 |
| Bone_Parietal_R | Anatomic | Bone | Parietal bone Right | 52788 |
| Bone_Sphenoid | Anatomic | Bone | Sphenoid Bone | 52736 |
| Bone_Temporal | Anatomic | Bone | Temporal Bone | 52737 |
| Bone_Temporal_L | Anatomic | Bone | Temporal Bone Left | 52739 |
| Bone_Temporal_R | Anatomic | Brain | Temporal Bone Right | 52738 |
| Bone_Zygomatic_L | Anatomic | Bone | Zygomatic Bone Left | 52893 |
| Bone_Zygomatic_R | Anatomic | Bone | Zygomatic Bone Right | 52892 |
| Bone_Zygomatics | Anatomic | Bone | Zygomatic Bone | 52747 |
| Brain | Anatomic | Brain | Brain | 50801 |
| Brain-CTV | Derived | Brain | Brain minus the CTV | — |
| Brain-GTV | Derived | Brain | Brain minus the GTV | — |
| Brain-PTV | Derived | Brain | Brain minus the PTV | — |
| Brainstem | Anatomic | Nerve | Brain Stem | 79876 |
| Brainstem_Core | Anatomic | Nerve | Core of the brainstem | — |
| Brainstem_PRV | PRV | Nerve | PRV for the Brainstem | — |
| Brainstem_PRVxx | PRV | Nerve | PRV margin on the brain stem that is an xx millimeter expansion | — |
| Brainstem_Surf | Anatomic | Nerve | Surface of the brainstem | — |
| Cavity_Nasal | Anatomic | Nose | Nasal Cavity | 54378 |
| Cavity_Oral | Anatomic | Mouth | Oral cavity | 20292 |
| Cerebellum | Anatomic | Brain | Cerebellum | 67944 |
| Cerebrum | Anatomic | Brain | Cerebrum | 62000 |
| Cerebrum_L | Anatomic | Brain | 61819 | |
| Cerebrum_R | Anatomic | Brain | 67292 | |
| Cist_Pontine | Anatomic | Brain | Pontine Cistern | 83719 |
| Cist_Suprasellar | Anatomic | Brain | — | |
| CN_III | Anatomic | Nerve | Third Cranial Nerve (Oculomotor nerve) | 50864 |
| CN_III_L | Anatomic | Nerve | Third Cranial Nerve (Oculomotor nerve) Left | 50880 |
| CN_III_R | Anatomic | Nerve | Third Cranial Nerve (Oculomotor nerve) Right | 50879 |
| CN_IX | Anatomic | Nerve | Ninth Cranial Nerve (Glossopharyngeal nerve) | 50870 |
| CN_IX_L | Anatomic | Nerve | Ninth Cranial Nerve (Glossopharyngeal nerve) Left | 50892 |
| CN_IX_R | Anatomic | Nerve | Ninth Cranial Nerve (Glossopharyngeal nerve) Right | 50870 |
| CN_V | Anatomic | Nerve | Fifth Cranial Nerve (Trigeminal nerve) | 50866 |
| CN_V_L | Anatomic | Nerve | Fifth Cranial Nerve (Trigeminal nerve) Left | 50885 |
| CN_V_R | Anatomic | Nerve | Fifth Cranial Nerve (Trigeminal nerve) Right | 50884 |
| CN_VI | Anatomic | Nerve | Sixth Cranial Nerve (Abducens nerve) | 50867 |
| CN_VI_L | Anatomic | Nerve | Sixth Cranial Nerve (Abducens nerve) Left | 50887 |
| CN_VI_R | Anatomic | Nerve | Sixth Cranial Nerve (Abducens nerve) Right | 50886 |
| CN_VII | Anatomic | Nerve | Seventh Cranial Nerve (Facial) | 50868 |
| CN_VII_L | Anatomic | Nerve | Seventh Cranial Nerve (Facial) Left | 50889 |
| CN_VII_R | Anatomic | Nerve | Seventh Cranial Nerve (Facial) Right | 50888 |
| CN_VIII | Anatomic | Nerve | Eighth Cranial (Vestibulocochlear) Nerve | 50869 |
| CN_VIII_L | Anatomic | Nerve | Eighth Cranial (Vestibulocochlear) Nerve Left | 50891 |
| CN_VIII_R | Anatomic | Nerve | Eighth Cranial (Vestibulocochlear) Nerve Right | 50890 |
| CN_XI | Anatomic | Nerve | Eleventh Cranial Nerve (Spinal accessory nerve) | 6720 |
| CN_XI_L | Anatomic | Nerve | Eleventh Cranial Nerve (Spinal accessory nerve) Left | 50899 |
| CN_XI_R | Anatomic | Nerve | Eleventh Cranial Nerve (Spinal accessory nerve) Right | 50897 |
| CN_XII | Anatomic | Nerve | Twelfth Cranial Nerve (Hypoglossal nerve) | 50871 |
| CN_XII_L | Anatomic | Nerve | Twelfth Cranial Nerve (Hypoglossal nerve) Left | 50903 |
| CN_XII_R | Anatomic | Nerve | Twelfth Cranial Nerve (Hypoglossal nerve) Right | 50901 |
| Cochlea | Anatomic | Ear | Cochlea | 60201 |
| Cochlea_L | Anatomic | Ear | Left Cochlea | 60203 |
| Cochlea_R | Anatomic | Ear | Right Cochlea | 60202 |
| Cornea | Anatomic | Eye | Cornea | 58238 |
| Cornea_L | Anatomic | Eye | Cornea Left | 58240 |
| Cornea_R | Anatomic | Eye | Cornea Right | 58239 |
| CribriformPlate | Anatomic | Bone | Cribriform Plate | 52890 |
| Cricoid | Anatomic | Cartilage | Cricoid cartilage | 9615 |
| Cricopharyngeus | Anatomic | Pharynx | Cricopharyngeal part of inferior pharyngeal constrictor | 46661 |
| Dens | Anatomic | Bone | Cervical vertebrae – Bony part of dens of axis | 24043 |
| Ear_External_L | Anatomic | Ear | External Ear Left | 53644 |
| Ear_External_R | Anatomic | Ear | External Ear Right | 53643 |
| Ear_Externals | Anatomic | Ear | External Ear | 52781 |
| Ear_Internal_L | Anatomic | Ear | Internal Ear Left | 61021 |
| Ear_Internal_R | Anatomic | Ear | Internal Ear Right | 61020 |
| Ear_Internals | Anatomic | Ear | Internal Ear | 60909 |
| Ear_Middle | Anatomic | Ear | Middle Ear | 56513 |
| Ear_Middle_L | Anatomic | Ear | Middle Ear Left | 56515 |
| Ear_Middle_R | Anatomic | Ear | Middle Ear Right | 56514 |
| Eye_L | Anatomic | Eye | Eyeball Left | 12515 |
| Eye_R | Anatomic | Eye | Eyeball Right | 12514 |
| Eyes | Anatomic | Eye | Set of eyes | 268861 |
| Fossa_Jugular | Anatomic | Bone | Jugular Fossa | 56429 |
| Fossa_Posterior | Anatomic | Bone | Posterior Fossa | 54368 |
| Glnd_Lacrimal | Anatomic | Gland | Lacrimal Gland | 59101 |
| Glnd_Lacrimal_L | Anatomic | Gland | Lacrimal Gland Left | 59103 |
| Glnd_Lacrimal_R | Anatomic | Gland | Lacrimal Gland Right | 59102 |
| Glnd_Subling_L | Anatomic | Gland | Sublingual gland | 59805 |
| Glnd_Subling_R | Anatomic | Gland | Sublingual gland | 59804 |
| Glnd_Sublings | Anatomic | Gland | Sublingual gland | 320440 |
| Glnd_Submand_L | Anatomic | Gland | Submandibular Gland Left | 59803 |
| Glnd_Submand_R | Anatomic | Gland | Submandibular Gland Right | 59802 |
| Glnd_Submands | Anatomic | Gland | Submandibular Gland | 320442 |
| Glottis | Anatomic | Glottis | Glottis | 55414 |
| Hardpalate | Anatomic | Head | Hard palate | 55023 |
| Hemisphere_L | Anatomic | Brain | 61819 | |
| Hemisphere_R | Anatomic | Brain | 67292 | |
| Hemispheres | Anatomic | Brain | — | |
| Hippocampi | Anatomic | Brain | Hippocampus | 275020 |
| Hippocampus_L | Anatomic | Brain | Hippocampus Left | 275024 |
| Hippocampus_R | Anatomic | Brain | Hippocampus Right | 275022 |
| Hypothalmus | Anatomic | Brain | Hypothalamus | 62008 |
| Hypothalmus_PRV | PRV | Brain | — | |
| Hypothalmus_PRVx | PRV | Brain | — | |
| Joint_TM | Anatomic | Joint | Temperomandibular Joint | 54832 |
| Joint_TM_L | Anatomic | Joint | Temperomandibular Joint Left | 54834 |
| Joint_TM_R | Anatomic | Joint | Temperomandibular Joint Right | 54833 |
| Laryngl_Pharynx | Anatomic | Tissue | Laryngeal pharynx | 54880 |
| Larynx | Anatomic | Larynx | Larynx | 55097 |
| Larynx_SG | Anatomic | Larynx | Supraglottic Larynx | 55476 |
| Lens | Anatomic | Eye | Eye Lens | 58241 |
| Lens_L | Anatomic | Eye | Lens Left | 58243 |
| Lens_R | Anatomic | Eye | Lens Right | 58242 |
| Lips | Anatomic | Feature | Lips | 59815 |
| LN_Neck_IA_L | Anatomic | Lymph Node | Level IA (Submental) neck node Left | 235616 |
| LN_Neck_IA_R | Anatomic | Lymph Node | Level IA (Submental) neck node Right | 235614 |
| LN_Neck_IB_L | Anatomic | Lymph Node | Level IB (Submandibular) neck node Left | 232676 |
| LN_Neck_IB_R | Anatomic | Lymph Node | Level IB (Submandibular) neck node Right | 232673 |
| LN_Neck_II_L | Anatomic | Lymph Node | Level IIA & IIB (Upper Jugular) neck nodes Left | 265660 |
| LN_Neck_II_R | Anatomic | Lymph Node | Level IIA & IIB (Upper Jugular) neck nodes Left | 265658 |
| LN_Neck_IIA_L | Anatomic | Lymph Node | Level IIA (Upper Jugular) neck node Left | 241975 |
| LN_Neck_IIA_R | Anatomic | Lymph Node | Level IIA (Upper Jugular) neck node Right | 241973 |
| LN_Neck_IIB_L | Anatomic | Lymph Node | Level IIB (Upper Jugular) neck node Left | 241979 |
| LN_Neck_IIB_R | Anatomic | Lymph Node | Level IIB (Upper Jugular) neck node Right | 241977 |
| LN_Neck_III_L | Anatomic | Lymph Node | Level III (Middle Jugular) neck node Left | 241953 |
| LN_Neck_III_R | Anatomic | Lymph Node | Level III (Middle Jugular) neck node Right | 241951 |
| LN_Neck_IV_L | Anatomic | Lymph Node | Level IV neck (Lower Jugular) node Left | 241959 |
| LN_Neck_IV_R | Anatomic | Lymph Node | Level IV (Lower Jugular) neck node Right | 241957 |
| LN_Neck_V_L | Anatomic | Lymph Node | Level VA, VB and VC (Posterior Triangle) neck nodes Left | 241965 |
| LN_Neck_V_R | Anatomic | Lymph Node | Level VA, VB and VC (Posterior Triangle) neck nodes Right | 241963 |
| LN_Neck_VA_L | Anatomic | Lymph Node | Level VA (Posterior Triangle) neck node Left | 265629 |
| LN_Neck_VA_R | Anatomic | Lymph Node | Level VA (Posterior Triangle) neck node Right | 265626 |
| LN_Neck_VB_L | Anatomic | Lymph Node | Level VB (Posterior Triangle) neck node Left | — |
| LN_Neck_VB_R | Anatomic | Lymph Node | Level VB (Posterior Triangle) neck node Right | — |
| LN_Neck_VC_L | Anatomic | Lymph Node | Level VC (Posterior Triangle) neck node Left | 232721 |
| LN_Neck_VC_R | Anatomic | Lymph Node | Level VC (Posterior Triangle) neck node Right | 232719 |
| LN_Neck_VI_L | Anatomic | Lymph Node | Level VI (Anterior Triangle) neck node Left | 241971 |
| LN_Neck_VI_R | Anatomic | Lymph Node | Level VI (Anterior Triangle) neck node Right | 241969 |
| LN_Neck_VII_L | Anatomic | Lymph Node | Level VII (Upper Mediastinal) neck node Left | — |
| LN_Neck_VII_R | Anatomic | Lymph Node | Level VII (Upper Mediastinal) neck node Right | — |
| Lobe_Frontal | Anatomic | Brain | Frontal Lobe | 61824 |
| Lobe_Frontal_L | Anatomic | Brain | Frontal Lobe Left | 72970 |
| Lobe_Frontal_R | Anatomic | Brain | Frontal Lobe Left | 72969 |
| Lobe_Occipital | Anatomic | Brain | Occipital Lobe | 67325 |
| Lobe_Occipital_L | Anatomic | Brain | Occipital Lobe Left | 72976 |
| Lobe_Occipital_R | Anatomic | Brain | Occipital Lobe Right | 72975 |
| Lobe_Parietal | Anatomic | Brain | Parietal Lobe | 61826 |
| Lobe_Parietal_L | Anatomic | Brain | Parietal Lobe Left | 72974 |
| Lobe_Parietal_R | Anatomic | Brain | Parietal Lobe Right | 72973 |
| Lobe_Temporal | Anatomic | Brain | Temporal Lobe | 61825 |
| Lobe_Temporal_L | Anatomic | Brain | Temporal Lobe Left | 72972 |
| Lobe_Temporal_R | Anatomic | Brain | Temporal Lobe Right | 72971 |
| Malleus | Anatomic | Bone | Malleus | 52753 |
| Malleus_L | Anatomic | Bone | Malleus Left | 74053 |
| Malleus_R | Anatomic | Bone | Malleus Right | 74052 |
| Maxilla | Anatomic | Bone | Maxilla | 9711 |
| Maxilla_L | Anatomic | Bone | Maxilla Left | 53650 |
| Maxilla_R | Anatomic | Bone | Maxilla Right | 53649 |
| Musc_Constrict | Anatomic | Muscle | Constrictor Muscle of Pharynx | 46620 |
| Musc_Constrict_I | Anatomic | Muscle | Pharynx – Inferior pharyngeal constrictor | 46623 |
| Musc_Constrict_M | Anatomic | Muscle | Pharynx – Middle pharyngeal constrictor | 46622 |
| Musc_Constrict_S | Anatomic | Muscle | Pharynx – Superior pharyngeal constrictor | 46621 |
| Musc_Masseter | Anatomic | Muscle | Masseter Muscle | 48996 |
| Musc_Masseter_L | Anatomic | Muscle | Masseter Muscle Left | 48998 |
| Musc_Masseter_R | Anatomic | Muscle | Masseter Muscle Right | 48997 |
| Musc_Platysma_L | Anatomic | Muscle | Platysma Left | 45740 |
| Musc_Platysma_R | Anatomic | Muscle | Platysma Right | 45739 |
| Musc_Pterygoid_L | Anatomic | Muscle | Pterygoid muscles – Left | — |
| Musc_Pterygoid_R | Anatomic | Muscle | Pterygoid muscles – Right | — |
| Musc_Sclmast_L | Anatomic | Muscle | Sternocleidomastoid Left | 13409 |
| Musc_Sclmast_R | Anatomic | Muscle | Sternocleidomastoid Left | 13408 |
| Musc_Temporal_L | Anatomic | Muscle | Temporal muscle – Left | 49008 |
| Musc_Temporal_R | Anatomic | Muscle | Temporal muscle – Right | 49007 |
| Nasalconcha_LI | Anatomic | Nose | Inferior Nasal Concha Left | 54738 |
| Nasalconcha_RI | Anatomic | Nose | Inferior Nasal Concha Right | 54737 |
| Nasopharynx | Anatomic | Nose | Nasal part of pharynx | 54878 |
| Nose | Anatomic | Nose | Nose | 46472 |
| OpticChiasm | Anatomic | Nerve | Optic chiasm | 62045 |
| OpticChiasm_PRV | PRV | Nerve | PRV for Optic Chiasm | — |
| OpticChiasm_PRVx | PRV | Nerve | PRV expansion of x mm on the optic chiasm | — |
| OpticNrv | Anatomic | Nerve | Optic nerve | 50863 |
| OpticNrv_L | Anatomic | Nerve | Optic nerve | 50878 |
| OpticNrv_PRV | PRV | Nerve | PRV for Optic Nerve | — |
| OpticNrv_PRV_L | PRV | Nerve | — | |
| OpticNrv_PRV_R | PRV | Nerve | — | |
| OpticNrv_PRVxx_L | PRV | Nerve | PRV expansion of xx millimeters on the right optic nerve | — |
| OpticNrv_PRVxx_R | PRV | Nerve | PRV created with xx mm expansion on the left optic nerve | — |
| OpticNrv_R | Anatomic | Nerve | Optic nerve | 50875 |
| Orbit_L | Anatomic | Eye | Orbit Left | 54668 |
| Orbit_R | Anatomic | Eye | Orbit Right | 54667 |
| Oropharynx | Anatomic | Throat | Oral part of pharynx | 54879 |
| Palate_Soft | Anatomic | Muscle | Soft palate | 55021 |
| Parotid_L | Anatomic | Gland | Parotid Left | 59798 |
| Parotid_R | Anatomic | Gland | Parotid Right | 59797 |
| Parotids | Anatomic | Gland | Pair of Parotid Glands | 320436 |
| Pharynx | Anatomic | Throat | Pharynx | 46688 |
| Pineal | Anatomic | Gland | Pineal gland | 62033 |
| Pituitary | Anatomic | Gland | Pituitary gland | 13889 |
| Pituitary_PRVxx | PRV | Gland | PRV constructed as xx millimeter expansion on Pituitary | — |
| Pons | Anatomic | Brain | Pons | 67943 |
| Proc_Condyloid_L | Anatomic | Bone | Condyloid process of mandible – Right | 52838 |
| Proc_Condyloid_R | Anatomic | Bone | Condyloid process of mandible – Left | 52840 |
| Proc_Coronoid_L | Anatomic | Bone | Coronoid process of mandible – Left | 52835 |
| Proc_Coronoid_R | Anatomic | Bone | Coronoid process of mandible – Right | 52834 |
| Pterygoid_Lat_L | Anatomic | Muscle | Pterygoid muscles lateral – Left | 49017 |
| Pterygoid_Lat_R | Anatomic | Muscle | Pterygoid muscles lateral – Right | 49016 |
| Pterygoid_Med_L | Anatomic | Muscle | Pterygoid muscles medial – Left | 49013 |
| Pterygoid_Med_R | Anatomic | Muscle | Pterygoid muscles medial – Right | 49012 |
| Retina_L | Anatomic | Eye | Retina Left | 58303 |
| Retina_PRVxx_L | PRV | Eye | — | |
| Retina_PRVxx_R | PRV | Eye | PRV constructed as xx millimeter expansion on Retina | — |
| Retina_R | Anatomic | Eye | Retina Right | 58302 |
| Retinas | Anatomic | Eye | Both Retinas | 58301 |
| Scalp | Anatomic | Skin | 46494 | |
| Sinus_Ethmoid | Anatomic | Sinus | Ethmoidal Sinus | 84115 |
| Sinus_Frontal | Anatomic | Brain | Frontal Sinus | 57417 |
| Sinus_Frontal_L | Anatomic | Brain | Frontal Sinus Left | 57419 |
| Sinus_Frontal_R | Anatomic | Brain | Frontal Sinus Right | 57418 |
| Sinus_Maxilry | Anatomic | Sinus | Maxillary Sinus | 57715 |
| Sinus_Maxilry_L | Anatomic | Sinus | Maxillary Sinus | 57717 |
| Sinus_Maxilry_R | Anatomic | Sinus | Maxillary Sinus | 57716 |
| Sinus_Sphenoid | Anatomic | Bone | Sphenoidal Sinus | 54683 |
| Sinus_Sphenoid_L | Anatomic | Bone | Sphenoidal Sinus Left | 54708 |
| Sinus_Sphenoid_R | Anatomic | Bone | Sphenoidal Sinus Right | 54707 |
| Skull | Anatomic | Bone | Skeletal system of head | 46565 |
| Spc_Retrophar_L | Anatomic | Space | Retropharyngeal space Left | — |
| Spc_Retrophar_R | Anatomic | Space | Retropharyngeal space Right | — |
| Spc_Retrophars | Anatomic | Space | Retropharyngeal space | 286702 |
| Spc_Retrosty | Anatomic | Space | Retrostyloid space | — |
| Spc_Retrosty_L | Anatomic | Space | Retrostyloid space -Left | — |
| Spc_Retrosty_R | Anatomic | Space | Retrostyloid space -Left | — |
| SpinalCord_Cerv | Anatomic | Nerve | Spinal Cord Cervical | 71166 |
| Stapes | Anatomic | Ear | Stapes | 52751 |
| Stapes_L | Anatomic | Ear | Stapes Left | 74049 |
| Stapes_R | Anatomic | Ear | Stapes Right | 74048 |
| Sys_Ventricular | Anatomic | Brain | 242787 | |
| Tongue | Anatomic | Mouth | Tongue | 54640 |
| Tongue_All | Anatomic | Mouth | — | |
| Tongue_Base | Anatomic | Mouth | 54645 | |
| Tongue_Base_L | Anatomic | Mouth | — | |
| Tongue_Base_R | Anatomic | Mouth | — | |
| Tongue_Oral | Anatomic | Mouth | 54644 | |
| Tongue_Oral_L | Anatomic | Mouth | 281502 | |
| Tongue_Oral_R | Anatomic | Mouth | 281500 | |
| Tonsil | Anatomic | Mouth | 9609 | |
| V_Jugular | Anatomic | Vein | Any Jugular Vein | — |
| V_Jugular_Ext_L | Anatomic | Vein | 13112 | |
| V_Jugular_Ext_R | Anatomic | Vein | 13111 | |
| V_Jugular_Int_L | Anatomic | Vein | Internal jugular vein Right | 4754 |
| V_Jugular_Int_R | Anatomic | Vein | Internal jugular vein Left | 4762 |
| VB_C | Anatomic | Bone | Cervical vertebrae | 72063 |
| VB_C1 | Anatomic | Bone | Atlas – C1 | 12519 |
| VB_C2 | Anatomic | Bone | Axis – C2 | 12520 |
| VB_C3 | Anatomic | Bone | Cervical vertebra – C3 | 12521 |
| VB_C4 | Anatomic | Bone | Cervical vertebra – C4 | 12522 |
| VB_C5 | Anatomic | Bone | Cervical vertebra – C5 | 12523 |
| VB_C6 | Anatomic | Bone | Cervical vertebra – C6 | 12524 |
| VB_C7 | Anatomic | Bone | Cervical vertebra – C7 | 12525 |
| VocalCord_L | Anatomic | Larynx | 55459 | |
| VocalCord_R | Anatomic | Larynx | 55458 | |
| VocalCords | Anatomic | Larynx | Vocal Cords | 323919 |
| Vomer | Anatomic | Bone | Vomer | 9710 |
Thorax — 180 structures
| TG-263 Name | Type | Category | Description | FMAID |
|---|---|---|---|---|
| A_Aorta | Anatomic | Artery | Aorta | 3734 |
| A_Aorta_Asc | Anatomic | Artery | Ascending Aorta | 3736 |
| A_Brachiocephls | Anatomic | Artery | Brachiocephalic Artery | 3932 |
| A_Coronary_L | Anatomic | Artery | Coronary Artery Left | 50040 |
| A_Coronary_R | Anatomic | Artery | Coronary Artery Right | 50039 |
| A_LAD | Anatomic | Artery | Anterior interventricular branch of LCA (left anterior descending artery) |
3862 |
| A_Pulmonary | Anatomic | Artery | Pulmonary Artery | 66326 |
| A_Subclavian | Anatomic | Artery | Subclavian Artery | 3951 |
| A_Subclavian_L | Anatomic | Artery | Subclavian Artery Left | 4694 |
| A_Subclavian_R | Anatomic | Artery | Subclavian Artery Right | 3953 |
| A_Vertebral | Anatomic | Artery | Vertebral arteries | 3956 |
| A_Vertebral_L | Anatomic | Artery | Vertebral arteries left | 4066 |
| A_Vertebral_R | Anatomic | Artery | Vertebral arteries right | 3958 |
| AirWay_Dist | Anatomic | Lung | Distal Airway | — |
| AirWay_Prox | Anatomic | Lung | Proximal Airway | — |
| Atrium | Anatomic | Heart | Atrium of the heart | 7099 |
| Atrium_L | Anatomic | Heart | Atrium of the heart Left | 7097 |
| Atrium_R | Anatomic | Heart | Atrium of the heart Right | 7096 |
| BrachialPlex_L | Anatomic | Nerve | Brachial plexus Left | 45245 |
| BrachialPlex_R | Anatomic | Nerve | Brachial plexus Right | 45244 |
| BrachialPlexs | Anatomic | Nerve | Brachial plexus | 5906 |
| Breast_L | Anatomic | Breast | Breast Left | 321497 |
| Breast_R | Anatomic | Breast | Breast Right | 321496 |
| Breasts | Anatomic | Breast | Both breasts | 268893 |
| Bronchus | Anatomic | Lung | Bronchial tree | 26660 |
| Bronchus_L | Anatomic | Lung | Bronchial tree Left | 26662 |
| Bronchus_Main | Anatomic | Lung | Main Bronchus | 7405 |
| Bronchus_Main_L | Anatomic | Lung | Main Bronchus Left | 7396 |
| Bronchus_Main_R | Anatomic | Lung | Main Bronchus Right | 7395 |
| Bronchus_PRVxx | Anatomic | Lung | A PRV expansion on the Bronchus that is xx millimeters thick | — |
| Bronchus_R | Anatomic | Lung | Bronchial tree Right | 26661 |
| Carina | Anatomic | Carina | Carina | 7465 |
| Cartlg_Thyroid | Anatomic | Cartilage | Thyroid cartilage | 55099 |
| Chestwall | Anatomic | Chest wall | Chest wall | 50060 |
| Chestwall_L | Anatomic | Chest wall | Left Chest Wall | 25559 |
| Chestwall_R | Anatomic | Chest wall | Right Chest Wall | 25558 |
| Clavicle_L | Anatomic | Bone | Clavicle Left | 13323 |
| Clavicle_R | Anatomic | Bone | Clavicle Right | 13322 |
| Diaphragm | Anatomic | Diaphragm | Diaphragm | 13295 |
| Esophagus | Anatomic | Esophagus | Esophagus | 7131 |
| Esophagus_I | Anatomic | Esophagus | Lower Esophagus (abdominal) | 9397 |
| Esophagus_M | Anatomic | Esophagus | Middle Esophagus (thoracic) | 9396 |
| Esophagus_NAdj | Anatomic | Esophagus | Non Adjacent Esophagus | — |
| Esophagus_S | Anatomic | Esophagus | Upper Esophagus (cervical) | — |
| Glnd_Adrenal_L | Anatomic | Gland | Adrenal glands left | 15629 |
| Glnd_Adrenal_R | Anatomic | Gland | Adrenal glands right | 15630 |
| Glnd_Parathyroid | Anatomic | Gland | Parathyroid gland | 13890 |
| Glnd_Thymus | Anatomic | Gland | Thymus Gland | 9607 |
| Glnd_Thyroid | Anatomic | Gland | Thyroid Gland | 9603 |
| GreatVes | Anatomic | Heart | Great Vessels of the heart (aorta, vena cava S&I, pulmonary A&V) | — |
| GreatVes_NAdj | Anatomic | Heart | Non Adjacent Great Vessels | — |
| Heart | Anatomic | Heart | Heart | 7088 |
| LN_Ax_Apical | Anatomic | Lymph Node | Set of apical axillary lymphatic vessels | 23394 |
| LN_Ax_Apical_L | Anatomic | Lymph Node | Axillary lymphatic chain – Apical Left | 73265 |
| LN_Ax_Apical_R | Anatomic | Lymph Node | Axillary lymphatic chain – Apical Right | 73264 |
| LN_Ax_Central_L | Anatomic | Lymph Node | Axillary lymphatic chain – Central Left | 73263 |
| LN_Ax_Central_R | Anatomic | Lymph Node | Axillary lymphatic chain – Central Left | 73262 |
| LN_Ax_Centrals | Anatomic | Lymph Node | Set of central axillary lymphatic vessels | 233482 |
| LN_Ax_L | Anatomic | Lymph Node | Axillary lymphatic chain Left | 73250 |
| LN_Ax_L1_L | Anatomic | Lymph Node | Level 1 Axillary Lymph Node Left | 276001 |
| LN_Ax_L1_R | Anatomic | Lymph Node | Level 1 Axillary Lymph Node Right | 275999 |
| LN_Ax_L2_L | Anatomic | Lymph Node | Level 2 Axillary Lymph Node Left | 276005 |
| LN_Ax_L2_R | Anatomic | Lymph Node | Level 2 Axillary Lymph Node Right | 276003 |
| LN_Ax_L3_L | Anatomic | Lymph Node | Level 3 Axillary Lymph Node Left | 276009 |
| LN_Ax_L3_R | Anatomic | Lymph Node | Level 3 Axillary Lymph Node Right | 276007 |
| LN_Ax_Lateral_L | Anatomic | Lymph Node | Axillary lymphatic chain – Lateral Left | 73256 |
| LN_Ax_Lateral_R | Anatomic | Lymph Node | Axillary lymphatic chain – Lateral Right | 73255 |
| LN_Ax_Laterals | Anatomic | Lymph Node | pair of Lungs | 233458 |
| LN_Ax_Pectoral_L | Anatomic | Lymph Node | Axillary lymphatic chain – Pectoral Left | 73253 |
| LN_Ax_Pectoral_R | Anatomic | Lymph Node | Axillary lymphatic chain – Pectoral Right | 73252 |
| LN_Ax_Pectorals | Anatomic | Lymph Node | Set of pectoral axillary lymphatic vessels | 233446 |
| LN_Ax_R | Anatomic | Lymph Node | Axillary lymphatic chain Right | 73249 |
| LN_Ax_Subscap_L | Anatomic | Lymph Node | Axillary lymphatic chain – Subscapular Left | 73259 |
| LN_Ax_Subscap_R | Anatomic | Lymph Node | Axillary lymphatic chain – Subscapular Right | 73258 |
| LN_Ax_Subscaps | Anatomic | Lymph Node | Set of subscapular axillary lymphatic vessels | 233470 |
| LN_Brachioceph_L | Anatomic | Lymph Node | Lymph nodes of thorax – Brachiocephalic Left | 5946 |
| LN_Brachioceph_R | Anatomic | Lymph Node | Lymph nodes of thorax – Brachiocephalic Right | 5945 |
| LN_Brachiocephs | Anatomic | Lymph Node | Lymph nodes of thorax – Brachiocephalic | 5944 |
| LN_Bronchpulm_L | Anatomic | Lymph Node | Lymph nodes of thorax – Bronchopulmonary Left | 5967 |
| LN_Bronchpulm_R | Anatomic | Lymph Node | Lymph nodes of thorax – Bronchopulmonary Right | 5966 |
| LN_Bronchpulms | Anatomic | Lymph Node | Lymph nodes of thorax – Bronchopulmonary | 5965 |
| LN_Diaphragmatic | Anatomic | Lymph Node | Lymph nodes of thorax – Diaphragmatic | 12773 |
| LN_IMN_L | Anatomic | Lymph Node | 5934 | |
| LN_IMN_R | Anatomic | Lymph Node | 5933 | |
| LN_IMNs | Anatomic | Lymph Node | Lymph nodes IMN | 5849 |
| LN_Intercostals | Anatomic | Lymph Node | Lymph nodes of thorax – Intercostal | 5932 |
| LN_Ligamentarter | Anatomic | Lymph Node | Lymph nodes of thorax – Ligamentum arteriosum | 74033 |
| LN_Mediastinals | Anatomic | Lymph Node | Lymph nodes of thorax – Mediastinal | 12774 |
| LN_Paramammary_L | Anatomic | Lymph Node | Lymph nodes of thorax – Paramammary Left | 232600 |
| LN_Paramammary_R | Anatomic | Lymph Node | Lymph nodes of thorax – Paramammary Right | 232598 |
| LN_Paramammarys | Anatomic | Lymph Node | Lymph nodes of thorax – Paramammary | 44313 |
| LN_Parasternal_L | Anatomic | Lymph Node | Lymph nodes of thorax – Parasternal Left | 5934 |
| LN_Parasternal_R | Anatomic | Lymph Node | Lymph nodes of thorax – Parasternal Right | 5933 |
| LN_Parasternals | Anatomic | Lymph Node | Lymph nodes of thorax – Parasternal | 5849 |
| LN_Pulmonary_L | Anatomic | Lymph Node | Lymph nodes of thorax – Pulmonary Left | 5970 |
| LN_Pulmonary_R | Anatomic | Lymph Node | Lymph nodes of thorax – Pulmonary Right | 5969 |
| LN_Pulmonarys | Anatomic | Lymph Node | Lymph nodes of thorax – Pulmonary | 5968 |
| LN_Supmammary_L | Anatomic | Lymph Node | Lymph nodes of thorax – Supramammary Left | 232604 |
| LN_Supmammary_R | Anatomic | Lymph Node | Lymph nodes of thorax – Supramammary Right | 232602 |
| LN_Supmammarys | Anatomic | Lymph Node | Lymph nodes of thorax – Supramammary | 12785 |
| LN_Trachbrnchs | Anatomic | Lymph Node | Lymph nodes of thorax – Tracheobronchial | 5950 |
| LN_Trachbrnchs_L | Anatomic | Lymph Node | Lymph nodes of thorax – Tracheobronchial Left | 5952 |
| LN_Trachbrnchs_R | Anatomic | Lymph Node | Lymph nodes of thorax – Tracheobronchial Right | 5951 |
| Lung_L | Anatomic | Lung | Lung Left | 7310 |
| Lung_LLL | Anatomic | Lung | Lung – lower lobe of left | 7371 |
| Lung_LUL | Anatomic | Lung | Lung – upper lobe of left | 7370 |
| Lung_R | Anatomic | Lung | Lung Right | 7309 |
| Lung_RLL | Anatomic | Lung | Lung – lower lobe of right | 7337 |
| Lung_RML | Anatomic | Lung | Lung – middle lobe of right | 7383 |
| Lung_RUL | Anatomic | Lung | Lung – upper lobe of right | 7333 |
| Lungs | Anatomic | Lung | Pair of Lungs | 68877 |
| Lungs-CTV | Derived | Lung | Total Lung minus the CTV | — |
| Lungs-GTV | Derived | Lung | Total Lung minus the GTV | — |
| Lungs-ITV | Derived | Lung | — | |
| Lungs-PTV | Derived | Lung | Total Lung minus the PTV | — |
| Mediastinum | Anatomic | Heart | Mediastinum | 9826 |
| Pacemaker | Non_Anatomic | Devices | Pacemaker | — |
| Pericardium | Anatomic | Heart | Pericardium | 9869 |
| Rib | Anatomic | Bone | Any Rib volume | 7574 |
| Rib01_L | Anatomic | Bone | First Rib Left | 7987 |
| Rib01_R | Anatomic | Bone | First Rib Right | 7857 |
| Rib02_L | Anatomic | Bone | Second rib Left | 8012 |
| Rib02_R | Anatomic | Bone | Second rib Right | 7882 |
| Rib03_L | Anatomic | Bone | Third rib Left | 8039 |
| Rib03_R | Anatomic | Bone | Third rib Right | 7909 |
| Rib04_L | Anatomic | Bone | Fourth rib Left | 8148 |
| Rib04_R | Anatomic | Bone | Fourth rib Right | 7957 |
| Rib05_L | Anatomic | Bone | Fifth rib Left | 8093 |
| Rib05_R | Anatomic | Bone | Fifth rib Right | 8066 |
| Rib06_L | Anatomic | Bone | Sixth rib Left | 8202 |
| Rib06_R | Anatomic | Bone | Sixth rib Right | 8175 |
| Rib07_L | Anatomic | Bone | Seventh rib Left | 8256 |
| Rib07_R | Anatomic | Bone | Seventh rib Right | 8229 |
| Rib08_L | Anatomic | Bone | Eighth rib Left | 8310 |
| Rib08_R | Anatomic | Bone | Eighth rib Right | 8283 |
| Rib09_L | Anatomic | Bone | Ninth rib Left | 8391 |
| Rib09_R | Anatomic | Bone | Ninth rib Right | 8364 |
| Rib10_L | Anatomic | Bone | Tenth rib Left | 8472 |
| Rib10_R | Anatomic | Bone | Tenth rib Right | 8445 |
| Rib11_L | Anatomic | Bone | Eleventh rib Left | 8532 |
| Rib11_R | Anatomic | Bone | Eleventh rib Right | 8531 |
| Rib12_L | Anatomic | Bone | Twelfth rib Left | 8534 |
| Rib12_R | Anatomic | Bone | Twelfth rib Right | 8533 |
| Scapula_L | Anatomic | Bone | Scapula Left | 13396 |
| Scapula_R | Anatomic | Bone | Scapula Right | 13395 |
| Spc_Supraclav_L | Anatomic | Space | Supraclavicular space – Left | 45797 |
| Spc_Supraclav_R | Anatomic | Space | Supraclavicular space – Right | 45796 |
| SpinalCord_Thor | Anatomic | Nerve | Spinal Cord Thoracic | 71167 |
| Thoracic_Duct | Anatomic | Duct | Thoracic Duct | 5031 |
| Trachea | Anatomic | Lung | Trachea | 7394 |
| Trachea_NAdj | Anatomic | Lung | Trachea non adjacent wall | — |
| V_Azygos | Anatomic | Vein | Azygos Vein | 4838 |
| V_Brachioceph_L | Anatomic | Vein | Brachiocephalic vein Left | 4761 |
| V_Brachioceph_R | Anatomic | Vein | Brachiocephalic vein Right | 4751 |
| V_Pulmonary | Anatomic | Vein | Pulmonary vein | 66643 |
| V_Subclavian_L | Anatomic | Vein | Subclavian Vein Left | 4763 |
| V_Subclavian_R | Anatomic | Vein | Subclavian Vein Right | 4755 |
| V_Subclavians | Anatomic | Vein | Subclavian Vein | 4725 |
| V_Venacava_I | Anatomic | Vein | Inferior vena cava | 10951 |
| V_Venacava_S | Anatomic | Vein | Superior vena cava | 4720 |
| Valve_Aortic | Anatomic | Valve | Aortic Valve | 7236 |
| Valve_Mitral | Anatomic | Valve | Mitral Valve | 7235 |
| Valve_Pulmonic | Anatomic | Valve | Pulmonic Valve | 7246 |
| Valve_Tricuspid | Anatomic | Valve | Tricuspid Valve | 7234 |
| VB_T | Anatomic | Bone | Thoracic Vertebra | 9139 |
| VB_T01 | Anatomic | Bone | Thoracic Vertebra T1 | 9165 |
| VB_T02 | Anatomic | Bone | Thoracic Vertebra T2 | 9187 |
| VB_T03 | Anatomic | Bone | Thoracic Vertebra T3 | 9209 |
| VB_T04 | Anatomic | Bone | Thoracic Vertebra T4 | 9248 |
| VB_T05 | Anatomic | Bone | Thoracic Vertebra T5 | 9922 |
| VB_T06 | Anatomic | Bone | Thoracic Vertebra T6 | 9945 |
| VB_T07 | Anatomic | Bone | Thoracic Vertebra T7 | 9968 |
| VB_T08 | Anatomic | Bone | Thoracic Vertebra T8 | 9991 |
| VB_T09 | Anatomic | Bone | Thoracic Vertebra T9 | 10014 |
| VB_T10 | Anatomic | Bone | Thoracic Vertebra T10 | 10037 |
| VB_T11 | Anatomic | Bone | Thoracic Vertebra T11 | 10059 |
| VB_T12 | Anatomic | Bone | Thoracic Vertebra T12 | 10081 |
| Ventricle | Anatomic | Heart | Ventricle (cardiac) | 7100 |
| Ventricle_L | Anatomic | Heart | Ventricle (cardiac) Left | 7101 |
| Ventricle_R | Anatomic | Heart | Ventricle (cardiac) Right | 7098 |
Abdomen — 60 structures
| TG-263 Name | Type | Category | Description | FMAID |
|---|---|---|---|---|
| A_Celiac | Anatomic | Artery | Celiac Artery | 50737 |
| A_Mesenteric_I | Anatomic | Artery | Inferior mesenteric artery | 14750 |
| A_Mesenteric_S | Anatomic | Artery | Superior mesenteric artery | 14749 |
| Appendix | Anatomic | Bowel | Appendix | 14542 |
| Bag_Bowel | Non_Anatomic | Bowel | Bowel Bag | — |
| BileDuct_Common | Anatomic | Bowel | Common bile duct | 14667 |
| Bowel | Anatomic | Bowel | 7199 | |
| Bowel_Small | Anatomic | Bowel | Small Bowel (small intestine) | 7200 |
| Cecum | Anatomic | Bowel | Large bowel – Cecum | 14541 |
| Colon | Anatomic | Bowel | Colon | 14543 |
| Colon_Ascending | Anatomic | Bowel | Large bowel – Ascending colon | 14545 |
| Colon_Decending | Anatomic | Bowel | Large bowel – Descending colon | 14547 |
| Colon_PTVxx | Anatomic | Bowel | PRV created with xx mm expansion on the left optic nerve | — |
| Colon_Sigmoid | Anatomic | Bowel | Large bowel – Sigmoid colon | 14548 |
| Colon_Transverse | Anatomic | Bowel | Large bowel -Transverse colon | 14546 |
| Duodenum | Anatomic | Gut | Small bowel – Duodenum | 7206 |
| Ileum | Anatomic | Gut | Small bowel – Ileum | 7208 |
| Jejunum | Anatomic | Gut | Small bowel – Jejunum | 7207 |
| Jejunum_Ileum | Anatomic | Gut | Both ileum and jejunum | — |
| Kidney_Cortex | Anatomic | Urinary | Renal cortex for both Kidneys | 15581 |
| Kidney_Cortex_L | Anatomic | Urinary | Renal cortex left | 15584 |
| Kidney_Cortex_R | Anatomic | Urinary | Renal cortex right | 15583 |
| Kidney_Hilum_L | Anatomic | Urinary | Renal Hilum Left | 15942 |
| Kidney_Hilum_R | Anatomic | Urinary | Renal Hilum Right | 15941 |
| Kidney_Hilums | Anatomic | Urinary | Renal Hilum for both Kidneys | 15610 |
| Kidney_L | Anatomic | Urinary | Kidney Left | 7205 |
| Kidney_L-GTV | Derived | Urinary | — | |
| Kidney_Pelvis_L | Anatomic | Urinary | Renal pelvis Left | 15579 |
| Kidney_Pelvis_R | Anatomic | Urinary | Renal pelvis Right | 15578 |
| Kidney_R | Anatomic | Urinary | Kidney Right | 7204 |
| Kidney_R-GTV | Derived | Urinary | — | |
| Kidney-GTV | Derived | Urinary | — | |
| Kidneys | Anatomic | Urinary | Both Kidneys | 264815 |
| Lig_Hepatogastrc | Anatomic | Ligament | Hepatogastric ligament | 16520 |
| Liver | Anatomic | Liver | Liver | 7197 |
| Liver-CTV | Derived | Liver | — | |
| Liver-GTV | Derived | Liver | Liver minus GTV | — |
| LN_Portahepatis | Anatomic | Lymph Node | Porta hepatis | 15758 |
| Musc_Digastric_L | Anatomic | Muscle | Digastric muscle Left | 46293 |
| Musc_Digastric_R | Anatomic | Muscle | Digastric muscle Right | 46292 |
| PancJejuno | Surgical | Gut | The pancreatic jejuno junction – generated by surgical procedure | — |
| Pancreas | Anatomic | Gland | Pancreas | 7198 |
| Pancreas_Head | Anatomic | Gland | Head of Pancreas | 10468 |
| Pancreas_Tail | Anatomic | Gland | Tail of Pancreas | 14519 |
| Peritoneum | Anatomic | Membrane | Peritoneal sac | 9908 |
| Skin_Peritoneum | Anatomic | Skin | — | |
| Spc_Bowel | Anatomic | Bowel | Space occupied by bowel | — |
| Spc_Bowel_Small | Anatomic | Space | — | |
| SpinalCord_Lum | Anatomic | Nerve | Spinal Cord Lumbar | 71168 |
| Spleen | Anatomic | Lymph Node | Spleen | 7196 |
| Spleen_Hilum | Anatomic | Lymph Node | Splenic hilum | 15841 |
| Stomach | Anatomic | Stomach | Stomach | 7148 |
| Stomach_PRVxx | PRV | Stomach | PRV created with xx mm expansion on the stomach | — |
| V_Portal | Anatomic | Vein | Portal Vein | 66645 |
| VB_L | Anatomic | Bone | Lumbar Vertebrae | 72065 |
| VB_L1 | Anatomic | Bone | Lumbar Vertebra L1 | 13072 |
| VB_L2 | Anatomic | Bone | Lumbar Vertebra L2 | 13073 |
| VB_L3 | Anatomic | Bone | Lumbar Vertebra L3 | 13074 |
| VB_L4 | Anatomic | Bone | Lumbar Vertebra L4 | 13075 |
| VB_L5 | Anatomic | Bone | Lumbar Vertebra L5 | 13076 |
Pelvis — 109 structures
| TG-263 Name | Type | Category | Description | FMAID |
|---|---|---|---|---|
| A_Iliac_Cflx_L | Anatomic | Artery | Circumflex Left Iliac Artery | — |
| A_Iliac_Cflx_R | Anatomic | Artery | Circumflex Right Iliac Artery | — |
| A_Iliac_Ext_L | Anatomic | Artery | External iliac artery Left | 18807 |
| A_Iliac_Ext_R | Anatomic | Artery | External iliac artery Right | 18806 |
| A_Iliac_Int_L | Anatomic | Artery | Internal iliac artery Left | 18810 |
| A_Iliac_Int_R | Anatomic | Artery | Internal iliac artery Right | 18809 |
| A_Iliac_L | Anatomic | Artery | Common iliac artery Left | 14766 |
| A_Iliac_R | Anatomic | Artery | Common iliac artery Right | 14765 |
| Acetabulum_L | Anatomic | Bone | Acetabulum | 16599 |
| Acetabulum_R | Anatomic | Bone | Acetabulum | 16598 |
| Acetabulums | Anatomic | Bone | Acetabulum | 16579 |
| Anus | Anatomic | Bowel | Anus | 15711 |
| Bladder | Anatomic | Bladder | Urinary Bladder | 15900 |
| Bladder_Wall | Anatomic | Bladder | Bladder Wall | 15902 |
| Bladder-CTV | Derived | Bladder | Bladder minus CTV | — |
| Bone_Ilium | Anatomic | Bone | Ilium | 42832 |
| Bone_Ilium_L | Anatomic | Bone | Ilium Left | 16591 |
| Bone_Ilium_R | Anatomic | Bone | Ilium Right | 16590 |
| Bone_Ischium_L | Anatomic | Bone | Ischium Left | 16594 |
| Bone_Ischium_R | Anatomic | Bone | Ischium Right | 16593 |
| Bone_Pelvic | Anatomic | Bone | Pelvic Bones (Bony Pelvis) | 16580 |
| Bone_Pelvic_L | Anatomic | Bone | Bony Pelvis Left | 20227 |
| Bone_Pelvic_R | Anatomic | Bone | Bony Pelvis Right | 20226 |
| Bowel_Large | Anatomic | Bowel | Large Bowel | 7201 |
| Canal_Anal | Anatomic | Bowel | Anal Canal | 15703 |
| CaudaEquina | Anatomic | Nerve | Cauda equina | 52590 |
| Cavernosum | Anatomic | Reproductive | Penis Corpus Cavernosum | 75189 |
| Cervix | Anatomic | Cervix | Cervix of uterus | 17740 |
| Femur_Base_L | Anatomic | Bone | Femur Base Left | 32846 |
| Femur_Base_R | Anatomic | Bone | Femur Base Right | 32845 |
| Femur_Head_L | Anatomic | Bone | Femur Head & Neck Left | 32843 |
| Femur_Head_R | Anatomic | Bone | Femur Head & Neck Right | 32842 |
| Femur_Joint_L | Anatomic | Joint | Femoral Joint Left | 35180 |
| Femur_Joint_R | Anatomic | Joint | Femoral Joint Right | 35179 |
| Femur_Neck_L | Anatomic | Bone | Femur Neck Right | 42386 |
| Femur_Neck_R | Anatomic | Bone | Femur Neck Left | 42387 |
| Femur_Shaft_L | Anatomic | Bone | Femur Shaft Left | 32849 |
| Femur_Shaft_R | Anatomic | Bone | Femur Shaft Right | 32848 |
| Foley | Non-Anatomic | Bladder | Foley Catheter | — |
| Gallbladder | Anatomic | Gallbladder | Gall bladder | 7202 |
| Genitals | Anatomic | Reproductive | Genitals | 45643 |
| LN_Iliac_Ext_L | Anatomic | Lymph Node | Lymph nodes of pelvis – external iliac Left | 229177 |
| LN_Iliac_Ext_R | Anatomic | Lymph Node | Lymph nodes of pelvis – external iliac Right | 229177 |
| LN_Iliac_Int_L | Anatomic | Lymph Node | Lymph nodes of pelvis – internal iliac Left | 224275 |
| LN_Iliac_L | Anatomic | Lymph Node | Lymph nodes of pelvis – common iliac Left | 224269 |
| LN_Iliac_R | Anatomic | Lymph Node | Lymph nodes of pelvis – common iliac Right | 224269 |
| LN_Inguinofem | Anatomic | Lymph Node | Lymph nodes of pelvis – inguinofemoral | 236337 |
| LN_Inguinofem_L | Anatomic | Lymph Node | Lymph nodes of pelvis – inguinofemoral | 236341 |
| LN_Inguinofem_R | Anatomic | Lymph Node | Lymph nodes of pelvis – inguinofemoral | 236339 |
| LN_lliac_Int_R | Anatomic | Lymph Node | Lymph nodes of pelvis – internal iliac Right | 224275 |
| LN_Obturator_L | Anatomic | Lymph Node | Lymph nodes of pelvis – obturator Left | 16676 |
| LN_Obturator_R | Anatomic | Lymph Node | Lymph nodes of pelvis – obturator Right | 16676 |
| LN_Paraaortic | Anatomic | Lymph Node | Lymph nodes of abdomen- para-aortic | 223899 |
| LN_Pelvic_L | Anatomic | Lymph Node | Pelvic Lymph Nodes Left | — |
| LN_Pelvic_R | Anatomic | Lymph Node | Pelvic Lymph Nodes Right | — |
| LN_Pelvics | Anatomic | Lymph Node | Pelvic Lymph Nodes | — |
| LN_Presacral_L | Anatomic | Lymph Node | Lymph nodes of pelvis – presacral Left | 234280 |
| LN_Presacral_R | Anatomic | Lymph Node | Lymph nodes of pelvis – presacral Right | 234280 |
| Ovaries | Anatomic | Reproductive | Ovary | 7409 |
| Ovary_L | Anatomic | Reproductive | Ovary Left | 7214 |
| Ovary_R | Anatomic | Reproductive | Ovary Right | 7213 |
| Parametrium | Anatomic | Reproductive | Parametrium | 77061 |
| PenileBulb | Anatomic | Reproductive | Penile Bulb | 19614 |
| Penis | Anatomic | Reproductive | Penis | 9707 |
| Perineum | Anatomic | Perineum | Perineum | 9579 |
| Prostate | Anatomic | Gland | Prostate | 9600 |
| ProstateBed | Anatomic | Surgery | Prostate Bed | — |
| PubicSymphys | Anatomic | Bone | Pubic Symphysis | 16950 |
| PubicSymphys_L | Anatomic | Bone | Pubic bone Left | 16597 |
| PubicSymphys_R | Anatomic | Bone | Pubic bone Right | 16596 |
| Rectal_Wall | Anatomic | Rectum | Rectal Wall | 14626 |
| Rectum | Anatomic | Rectum | Large bowel – Rectum | 14544 |
| SacralPlex | Anatomic | Nerve | Sacral Plexus | 5909 |
| Sacrum | Anatomic | Bone | Sacrum | 16202 |
| Scrotum | Anatomic | Skin | Scrotum (skin & cremasteric fascia) | 18252 |
| SeminalVes | Anatomic | Gland | Seminal vesicle | 19387 |
| SeminalVes_Dist | Anatomic | Gland | Distal Seminal Vesicle | — |
| SeminalVes_Prox | Anatomic | Gland | Proximal Seminal Vesicle | — |
| Skin_Perineum | Anatomic | Skin | 20429 | |
| Sphincter_Anal | Anatomic | Bowel | Internal Anal Sphincter | 15710 |
| SpinalCord_Sac | Anatomic | Nerve | Spinal Cord Sacral | 256623 |
| Spongiosum | Anatomic | Reproductive | Penis Corpus Spongiosum | 19617 |
| Testis | Anatomic | Reproductive | Testis | 7210 |
| Testis_L | Anatomic | Reproductive | Testis Left | 7212 |
| Testis_R | Anatomic | Reproductive | Testis Right | 7211 |
| Ureter_L | Anatomic | Urinary | Ureter Left | 17888 |
| Ureter_R | Anatomic | Urinary | Ureter Right | 17887 |
| UreterDivert | Anatomic | Urinary | Urinary Divergence | — |
| Ureters | Anatomic | Urinary | Both Ureters | 264814 |
| Urethra | Anatomic | Urinary | Urethra | 19667 |
| Urethra_Prostatc | Anatomic | Urinary | Prostatic Urethra | — |
| Uterus | Anatomic | Reproductive | Uterus | 17558 |
| V_Iliac_Ext_L | Anatomic | Vein | External iliac vein Left | 18886 |
| V_Iliac_Ext_R | Anatomic | Vein | External iliac vein Right | 18885 |
| V_Iliac_Int_L | Anatomic | Vein | Internal iliac vein Left | 18810 |
| V_Iliac_Int_R | Anatomic | Vein | Internal iliac vein Right | 18809 |
| V_Iliac_L | Anatomic | Vein | Common iliac vein Right | 21387 |
| V_Iliac_R | Anatomic | Vein | Common iliac vein Left | 21388 |
| Vagina | Anatomic | Reproductive | Vagina | 19949 |
| Vagina_Surf | Anatomic | Reproductive | — | |
| VaginalCuff | Anatomic | Reproductive | Vaginal Cuff | — |
| VB_S | Anatomic | Bone | Sacral Vertebra | 12526 |
| VB_S1 | Anatomic | Bone | Sacral Vertebra S1 | 13077 |
| VB_S2 | Anatomic | Bone | Sacral Vertebra S2 | 13078 |
| VB_S3 | Anatomic | Bone | Sacral Vertebra S3 | 13079 |
| VB_S4 | Anatomic | Bone | Sacral Vertebra S4 | 13080 |
| VB_S5 | Anatomic | Bone | Sacral Vertebra S5 | 13081 |
| Vulva | Anatomic | Reproductive | Vulva | 20462 |
| Wall_Vagina | Anatomic | Reproductive | Wall of vagina | 19971 |
Body / Systemic — 52 structures
| TG-263 Name | Type | Category | Description | FMAID |
|---|---|---|---|---|
| Bag_Ostomy | Non_Anatomic | Devices | Ostomy Bag | — |
| Body | Anatomic | Body | Only the body | 256135 |
| Body-PTV | Derived | Body | Body minus PTV | — |
| Bolus | Non_Anatomic | Bolus | — | |
| Bolus_xxmm | Non_Anatomic | Bolus | Bolus that is xx millimeters thick e.g. Bolus_03mm, Bolus_10mm | — |
| Bone | Anatomic | Bone | Bone | 30317 |
| BoneMarrow | Anatomic | Bone | Bone Marrow | 9608 |
| BoneMarrow_Act | Anatomic | Bone | Active Bone Marrow | — |
| Boost | Non-Anatomic | Body | Boost Volume | — |
| CTV | Target | CTV | Clinical Tumor Volume | — |
| Edema | Anatomic | Skin | Edema | — |
| E-PTV_Ev05_xxxx | Non-Anatomic | Body | All tissue excluding the 5 mm expanded PTV. Generated by subtracting the 5 mm expanded PTV receiving a dose of xxxx cGy from the external contour. | — |
| E-PTV_xxxx | Non-Anatomic | Derived | All tissue excluding the PTV. Generated by subtracting the PTV receiving a dose of xxxx cGy from the external contour. | — |
| Eval | Non-Anatomic | Body | Evaluation Structure | — |
| External | Anatomic | Body | Contour encompassing body plus other external items | — |
| GrowthPlate_L | Anatomic | Bone | — | |
| GrowthPlate_R | Anatomic | Bone | — | |
| GTV | Target | GTV | Gross Tumor Volume | — |
| IDL | Non-Anatomic | Dose | Isodose Line e.g. IDL_5000 isodose line for 50 Gy | — |
| ITV | Target | ITV | Internal Target Volume | — |
| Joint_Surface | Anatomic | Joint | — | |
| Leads | Non_Anatomic | Devices | — | |
| LN | Anatomic | Lymph Node | Lymph Node | 5034 |
| LN_L | Anatomic | Lymph Node | Lymph Node Left | — |
| LN_R | Anatomic | Lymph Node | Lymph Node Right | — |
| LN_Sclav_L | Anatomic | Lymph Node | Supraclavicular Lymph Node Left | 232719 |
| LN_Sclav_R | Anatomic | Lymph Node | Supraclavicular Lymph Node Right | 232721 |
| Markers | Non_Anatomic | Devices | — | |
| Musc | Anatomic | Muscle | Muscle | 30316 |
| Nrv_Peripheral | Anatomic | Nerve | Peripheral Nerve | — |
| Nrv_Root | Anatomic | Nerve | Nerve Root | 5981 |
| Postop | Non_Anatomic | Surgery | Post operative volume | — |
| Preop | Non_Anatomic | Surgery | Region originally occupied by the gross tumor volume | — |
| Prosthesis | Anatomic | Surgery | Prosthesis | — |
| PTV | Target | PTV | Planning Target Volume | — |
| Scar | Anatomic | Skin | Scar | — |
| Scar_Boost | Anatomic | Skin | — | |
| Skin | Anatomic | Skin | Skin | 7163 |
| Spc | Anatomic | Space | Space | — |
| SpinalCanal | Anatomic | Canal | Vertebral canal | 9680 |
| SpinalCanal_PRV | PRV | PRV | — | |
| SpinalCanal_PRVx | PRV | PRV | A PRV created with an x millimeter expansion on the spinal canal | — |
| SpinalCord | Anatomic | Nerve | Spinal Cord | 7647 |
| SpinalCord_PRV | PRV | PRV | — | |
| SpinalCord_PRVxx | PRV | PRV | A PRV created with an xx millimeter expansion on the spinal cord | — |
| Strct | Anatomic | Body | Structure | — |
| SurgicalBed | Anatomic | Surgery | — | |
| ThecalSac | Anatomic | Membrane | 83720 | |
| TumorBed | Non_Anatomic | Surgery | Tumor Bed | — |
| Valve | Anatomic | Valve | Valve | — |
| VB | Anatomic | Bone | Vertebral Body | — |
| VBs | Anatomic | Bone | Vertebral Bodies | 11945 |
Limb (singular) — 16 structures
| TG-263 Name | Type | Category | Description | FMAID |
|---|---|---|---|---|
| A_Femoral_Cflx_L | Anatomic | Artery | Circumflex Left Femoral Artery | — |
| A_Femoral_Cflx_R | Anatomic | Artery | Circumflex Right Femoral Artery | — |
| A_Femoral_L | Anatomic | Artery | Femoral Artery Left | 70250 |
| A_Femoral_R | Anatomic | Artery | Femoral Artery Right | 70249 |
| A_Humeral_Cflx_L | Anatomic | Artery | Circumflex Humeral Artery Left | — |
| A_Humeral_Cflx_R | Anatomic | Artery | Circumflex Humeral Artery Right | — |
| A_Humeral_L | Anatomic | Artery | Humeral Artery Left | — |
| A_Humeral_R | Anatomic | Artery | Humeral Artery Right | — |
| Femur_L | Anatomic | Bone | Femur Whole Left | 24475 |
| Femur_R | Anatomic | Bone | Femur Whole Right | 24474 |
| Femurs | Anatomic | Bone | Both Femurs | 9611 |
| Radius_L | Anatomic | Bone | Radius Left | 23465 |
| Radius_R | Anatomic | Bone | Radius Right | 23464 |
| Strct_Suprapatel | Anatomic | Joint | Suprapatellar Structures | — |
| Tendon | Anatomic | Tendon | 9721 | |
| Tendon_Quad | Anatomic | Tendon | 46900 |
Limbs (bilateral) — 17 structures
| TG-263 Name | Type | Category | Description | FMAID |
|---|---|---|---|---|
| Elbow | Anatomic | Joint | Elbow | 24901 |
| Elbow_L | Anatomic | Joint | Elbow Left | 24903 |
| Elbow_R | Anatomic | Joint | Elbow Right | 24902 |
| Fibula | Anatomic | Bone | Fibula | 24479 |
| Fibula_L | Anatomic | Bone | Fibula Left | 24481 |
| Fibula_R | Anatomic | Bone | Fibula Right | 24480 |
| Humerus_L | Anatomic | Bone | Humerus Left | 23131 |
| Humerus_R | Anatomic | Bone | Humerus Right | 23130 |
| Joint_Elbow | Anatomic | Joint | Elbow joint | 35289 |
| Joint_Elbow_L | Anatomic | Joint | Left Elbow joint | 35295 |
| Joint_Elbow_R | Anatomic | Joint | Right Elbow joint | 35294 |
| Joint_Glenohum | Anatomic | Joint | Glenohumeral Joint | 25912 |
| Joint_Glenohum_L | Anatomic | Joint | Glenohumeral Joint Left | 25929 |
| Joint_Glenohum_R | Anatomic | Joint | Glenohumeral Joint Right | 25927 |
| Knee | Anatomic | Joint | Knee | 24974 |
| Knee_L | Anatomic | Joint | Knee Left | 24978 |
| Knee_R | Anatomic | Joint | Knee Right | 24977 |
Integration with treatment planning systems and artificial intelligence
Nomenclature standardization gains operational relevance when it connects with the systems that consume and produce structure names on a daily basis: treatment planning systems (TPS), auto-segmentation tools, and data analysis platforms. TG-263 is not an isolated document — it is an interoperability layer that enables different links in the chain to communicate without manual mapping.
Why standardized nomenclature matters for AI
Deep learning-based auto-segmentation models depend on three conditions to perform well: large training datasets, consistent labels, and predictable mapping between the model output and the clinical structures in the treatment plan. TG-263 directly addresses the second and third conditions.
When a model is trained on contours labeled Parotid_L across all contributing institutions, the output label can be directly inserted into the plan without a translation step. If each institution uses its own name, the model needs a synonym dictionary that grows with every new dataset incorporated — and every missing entry in that dictionary is a structure that will not be correctly mapped in the clinic.
The scale of the problem becomes evident in projects like the TCIA (The Cancer Imaging Archive) and AAPM Grand Challenges, where data from dozens of institutions are combined for training and validation. Without standardized nomenclature, the harmonization step consumes more time than model training itself. With TG-263 adopted as the labeling convention, data arrives ready for use.
Tools that support TG-263 nomenclature
The radiation therapy ecosystem already offers tools that recognize, map, or enforce TG-263 nomenclature. The list below describes the main ones in an educational and neutral tone, without ranking.
AutoSeg (RT Medical Systems) is an auto-segmentation platform that produces contours with TG-263 names as its default output. Integration with RTConnect allows generated contours to flow directly to the TPS with compliant nomenclature, eliminating the manual renaming step. The complete pipeline — image reception, segmentation, naming, and delivery — respects the standard end to end.
Limbus Contour combines deep learning with atlas-based approaches for auto-contouring. The tool allows configuration of nomenclature templates that map model outputs to TG-263 names, facilitating integration with workflows that already follow the standard.
MVision AI operates as a cloud-based auto-contouring service, receiving images via DICOM and returning segmented structure sets. The system offers mapping of output names to TG-263 conventions, useful for departments that adopt the standard and receive contours from different providers.
Radformation AutoContour uses a combination of atlas and deep learning methods for automatic segmentation. The platform supports structure name customization, allowing departments to configure output to follow the TG-263 convention in their institutional template.
MIM Software offers contouring workflows with customizable structure templates. The ability to define templates with TG-263 names and apply them to new cases facilitates standard adoption in departments that process high patient volumes.
Varian Eclipse includes, starting from ARIA 13.x, a Structure Dictionary that associates TG-263 names with FMAIDs and recognized synonyms. When the physicist creates a new structure, the system suggests the standard name and allows synonym search. This feature lowers the adoption barrier because the professional does not need to memorize the entire catalog — the system provides guidance during delineation.
Elekta Monaco supports structure set management and templates that can be configured with TG-263 names. The template tool allows the department to predefine expected structures for each treatment site, enforcing nomenclature from the plan creation stage.
RaySearch RayStation offers scripting capabilities via IronPython and C# that allow building nomenclature validators. A script can check, upon saving the plan, whether all structure names adhere to the TG-263 standard and alert the user if deviations are found. This approach transforms nomenclature from a recommendation into an operational rule.
Impact on Big Data and multicenter research
The accumulated value of standardization becomes most apparent at scale. When dozens or hundreds of institutions feed a centralized repository with dosimetric and contour data — as occurs in Learning Health Systems, outcomes registries, and toxicity prediction projects — consistent nomenclature is a prerequisite for the data to be usable without intensive manual curation.
Projects that analyze correlations between organ-at-risk dose and clinical outcomes (NTCP models, for example) need to aggregate DVHs from thousands of patients. If each institution names the esophagus differently, the analysis pipeline requires an entity resolution step that is labor-intensive, error-prone, and difficult to audit. TG-263 eliminates this step by defining Esophagus as the canonical, universally recognized name.
The same principle applies to segmentation competitions such as the AAPM Grand Challenges and public repositories like the TCIA. Adoption of TG-263 as the labeling convention in these environments is already underway and is expected to become a formal requirement in future publications. Departments that adopt the standard now position their data to participate in these initiatives without retroactive rework.
Implementation workflow in 10 steps
Section 13 of the TG-263 report defines a 10-step implementation workflow designed to be applicable to any radiation therapy department, regardless of size or operational maturity. The workflow does not require halting operations — it is an incremental process that can be executed in parallel with clinical routine.
Step 1: Identify common treatment sites and involved staff. The starting point is mapping which disease sites the department treats most frequently and which professionals participate in delineation, planning, and review for each site. This mapping defines the initial scope of adoption and avoids the mistake of trying to standardize everything at once.
Step 2: Document current naming conventions. Before changing any name, the department needs to document what it already uses. This step reveals the true extent of inconsistency — it is common for teams to discover variations they did not know existed. Listing the names currently in use for the most frequent structures creates the comparison baseline for the TG-263 standard.
Step 3: Download the complete TG-263 list. The supplemental material from the report, available on the AAPM website, contains the spreadsheet with all 717 structures and all 9 columns. This spreadsheet is the reference document for the entire process.
Step 4: Save and create a working copy. The department should keep an intact copy of the original spreadsheet and work from a separate version. The working copy will be customized to the department’s reality; the original serves as an immutable reference for future consultation and for new professionals who need to understand the standard.
Step 5: Remove structures the department does not use. The complete spreadsheet contains 717 structures, but no department uses all of them. A center specializing in head and neck may start by removing pelvis and abdomen structures that are not part of its practice. This pruning reduces the catalog to a manageable size and focuses the team’s attention on the structures that actually matter.
Step 6: Discuss with disease-site groups and stakeholders. Each structure retained in the list should be validated by the professionals who use it. Physicians need to agree that the standard names are clinically appropriate. Physicists need to confirm that the names are compatible with existing scripts and tools. Dosimetrists need to verify that the names are practical in the daily delineation workflow.
Step 7: Identify templates and documents that need updating. Nomenclature does not live only in the TPS. Institutional protocols, QA checklists, prescription forms, automation scripts, and dosimetry reports may contain structure names. All need to be reviewed to reflect the adopted standard.
Step 8: Develop a phased rollout plan. The most effective implementation starts with one disease site (typically the most frequent), validates the standard with real cases, and only then expands to other sites. Each phase includes training, monitoring, and adjustment before moving forward. Attempts to implement all sites simultaneously typically result in partial adherence and eventual abandonment.
Step 9: Create TPS templates with standard names. Structure templates in the treatment planning system are the most effective enforcement mechanism. When the physicist or dosimetrist opens a new case and loads the “head and neck” template, all structures come pre-named according to TG-263. This eliminates manual typing and dramatically reduces the chance of deviation.
Step 10: Keep the complete list accessible for future reference. New professionals, new modalities, and new disease sites will emerge. The complete spreadsheet (AAPM TG-263 supplemental materials) should be available in a known location (network drive, intranet, document management system) so that anyone can consult the standard when they need to add a structure to the local catalog.
Relationship with target volume delineation
TG-263 nomenclature does not exist in isolation from the delineation process. On the contrary: it was designed to integrate directly into the contouring workflow, functioning as the identity layer that connects what the physician delineates to what the planning system calculates and what the database stores.
When a radiation oncologist delineates the GTV of a primary oropharyngeal tumor, TG-263 specifies that this structure should be named GTV_p. If there is nodal involvement, lymph nodes delineated as gross disease become GTV_n. The CTV covering the microscopic extension of the primary will be CTV_p, and the nodal one, CTV_n. The PTV generated from the primary CTV at the highest dose level will be PTV_p_High or PTV_7000 (if the dose is 70 Gy). Each name carries clinical information that any professional trained in the standard can decode without consulting the chart.
This integration is particularly valuable in head and neck sites, where the number of structures per plan can exceed 40 and the distinction between organs at risk, primary targets, and nodal targets must be unambiguous. A nasopharyngeal plan with three dose levels, multiple GTVs, and a dense network of organs at risk becomes significantly more readable when the nomenclature follows a predictable standard. The same clarity applies to other head and neck subsites: hypopharyngeal carcinoma, oral cavity, nasal and paranasal sinus tumors, salivary gland tumors, and thyroid cancer all benefit from unambiguous structure naming when plans involve overlapping organs at risk.
The same principle applies to all anatomic sites. In oropharyngeal delineation, the distinction between mucosal GTV and nodal GTV is immediately visible in the names. In larynx, where cartilage structures such as Arytenoid_L, Cricoid, and Thyroid_Cart are critical for margin definition, TG-263 names eliminate doubt about which cartilage is being referenced.
In breast treatments, standardized nomenclature facilitates the distinction between Breast_L (whole breast as an organ) and planning structures such as zBreast_Flash. Lung plans benefit from the clarity between Lung_L, Lung~_L (partial), and Lungs (bilateral), each name with distinct dosimetric implications. Regional nodal breast irradiation further demonstrates the value of TG-263, where consistent naming of supraclavicular, axillary, and internal mammary chain structures is essential for reproducible treatment planning.
In the pelvis, prostate delineation uses Prostate, SeminalVes, Rectum, Bladder, and Femur_L/Femur_R as standard organs at risk. For rectal cancer, nodal level nomenclature follows the TG-263 LN_ convention, making it easy to identify which lymph node chains are included in the CTV. Bladder and gynecologic delineation follow the same logic. Additional pelvic sites such as anal cancer, postoperative gynecologic radiotherapy, vulvar cancer, and testicular seminoma equally rely on consistent naming for pelvic organs at risk and nodal chains.
In scenarios such as brain metastases treated with SRS, target volume nomenclature with number identification (GTV_1, GTV_2, PTV_1) becomes critical for traceability. TG-263 allows this numbering within the name composition framework. For intracranial lesions, whether benign CNS tumors such as meningiomas and vestibular schwannomas or malignant CNS tumors such as high-grade gliomas, standardized naming of critical structures like Brainstem, Hippocampus_L, and OpticChiasm ensures reliable plan comparison and outcomes analysis.
The connection between nomenclature and delineation extends to special techniques as well. In image-guided brachytherapy, structures such as Applicator, Tandem, and pelvic organs at risk follow the same standard. For esophageal cancer, lymphoma, and soft tissue sarcoma, TG-263 provides standard names for the site-specific anatomical structures. In abdominal sites, gastric cancer, pancreatic cancer, and hepatocellular carcinoma delineation all benefit from consistent naming of structures such as Stomach, Duodenum, Liver, and Kidney_L/Kidney_R, enabling automated constraint checking and cross-institutional dose comparison.
For a comprehensive overview of how target volume delineation integrates into the radiation therapy workflow, see our complete target volume delineation guide, which covers principles, techniques, and recommendations by anatomic site.
The future of standardization in radiation therapy
TG-263 represents the state of the art in structure nomenclature for radiation therapy, but the field continues to evolve. Several developments underway promise to expand the reach and depth of standardization.
Integration with DICOM RT Structure Set. The DICOM standard already supports the concept of a Structure Set, but adoption of TG-263 nomenclature as a controlled vocabulary within DICOM is under discussion. Supplement 196, proposed by DICOM Working Group 7, defines a mechanism for including structure templates directly in the DICOM standard, enabling systems to interoperate not only on data format but also on the semantics of names. When this supplement is widely implemented, the name Parotid_L in a DICOM file will be not just a label, but a semantic identifier linked to an ontology.
HL7 FHIR and medical record interoperability. The HL7 FHIR standard, increasingly adopted for clinical information exchange, offers resources for coding procedures and anatomical findings. The bridge between TG-263 (radiation therapy nomenclature) and FHIR (health interoperability) runs through the mapping of FMAIDs and SNOMED CT codes. This mapping allows radiation therapy data to be integrated into the electronic health record with preserved semantics, enabling queries such as “which patients received dose > 50 Gy to the left parotid” directly in the hospital information system.
AI model registries. As auto-segmentation models proliferate, the need arises for registries that document the training nomenclature of each model. If a model was trained with TG-263 labels, any department that uses the standard can adopt it without mapping. If it was trained with proprietary labels, the mapping needs to be documented and validated. The trend is for model registries to include the TG-263 mapping table as mandatory metadata.
Multilingual translation tables. TG-263 was published in English, but departments in non-English-speaking countries need mapping between TG-263 names and local names. Translation tables associating, for example, Brainstem with “tronco encefalico” and SpinalCord with “medula espinhal” are already in development in several countries. The expectation is that the AAPM or an international working group will publish official correspondence tables that preserve the TG-263 name as the canonical identifier and offer translations as interface aliases.
Task group succession. The original TG-263 was published in 2018 and covers anatomical structures known up to that date. New treatment techniques, new imaging modalities, and new delineation approaches will continue to demand additions to the catalog. Maintaining the standard will require a successor group — whether linked to the AAPM, ASTRO, or a joint entity — that periodically reviews the spreadsheet, incorporates new structures, and publishes updated versions. The governance model of this group will determine whether TG-263 remains a living standard or freezes at the 2018 version.
Conclusion
TG-263 transformed structure nomenclature in radiation therapy from a matter of personal preference into a documented, tested, and adoptable technical standard. The 717 cataloged structures, the 15 principles for organs at risk, the 10 principles for target volumes, and the standardized DVH notation form a complete toolkit for any department that wants to operate with consistency.
The clinical impact is direct. Faster plan review. Usable historical data. Reliable QA automation. Compatibility with auto-segmentation models. Participation in multicenter studies without rework. Easier training of new professionals. None of these benefits depends on sophisticated technology — it depends on naming discipline.
Implementation does not need to be a months-long project. The report’s own 10-step workflow shows that starting with one disease site, creating TPS templates, and training the team already produces tangible results in the first week. Expansion to other sites happens naturally as the team internalizes the logic of the standard.
If your department has not yet adopted TG-263, the best time to start is now. Download the spreadsheet, identify the structures for your most frequent treatment site, and create the first standardized template. RT Medical Systems offers tools such as AutoSeg and RTConnect that integrate TG-263 nomenclature directly into the clinical workflow — from automatic segmentation to contour delivery in the TPS, with compliant names from the start. If you need support on this journey, our team is available to assess your department’s scenario and propose a customized adoption plan.