VoxTell is a 3D vision-language model that segments structures in volumetric medical images from free-text prompts. Trained on over 62,000 CT, MRI, and PET volumes covering more than 1,000 anatomical and pathological classes, the model represents a concrete advance in automatic segmentation. Open source integrations connect VoxTell to interactive web interfaces and to the Varian Eclipse ESAPI, creating research prototypes that bring academic models closer to the actual radiotherapy workflow.
This article details the model architecture, both public integrations — web and ESAPI — and the DICOM coordinate conversion pipeline that makes it all possible. All content refers exclusively to research and technical evaluation tools, never to clinical software.
What VoxTell Changes in 3D Segmentation
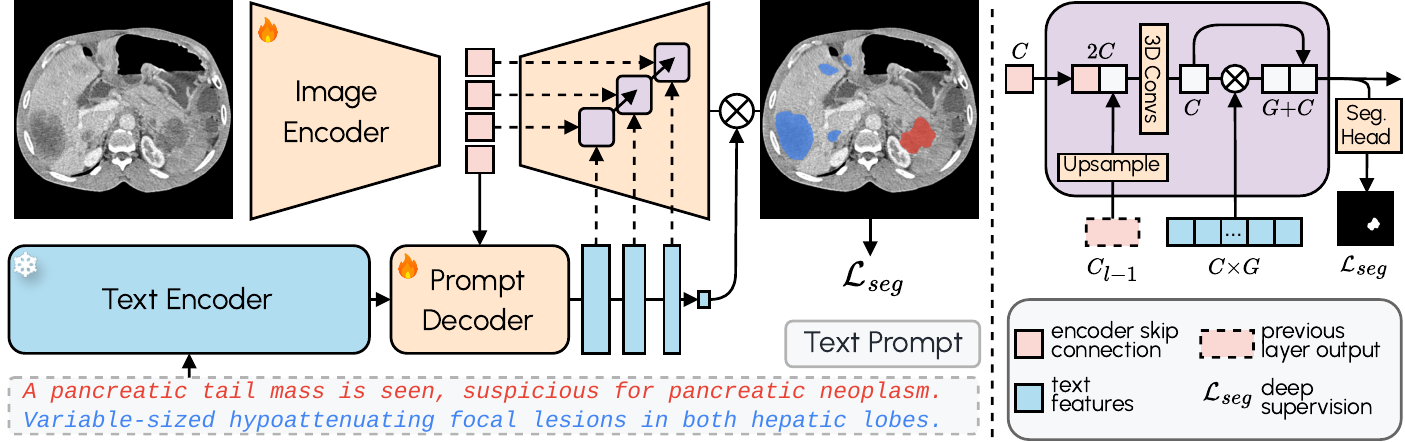
Conventional segmentation models work with fixed labels. If the model was not trained for “posterior fossa tumor,” it simply cannot segment it. VoxTell replaces this paradigm with free-text prompts: the operator types the desired structure — from “liver” to “left kidney with cortical cyst” — and the model generates the corresponding volumetric mask.
The architecture combines a 3D image encoder with Qwen3-Embedding-4B as a frozen text encoder. A prompt decoder transforms textual queries and latent image representations into multi-scale text features. The image decoder fuses visual and textual information at multiple resolutions using MaskFormer-style query-image fusion with deep supervision. The result: zero-shot segmentation with state-of-the-art performance on familiar structures and reasonable generalization to unseen classes.
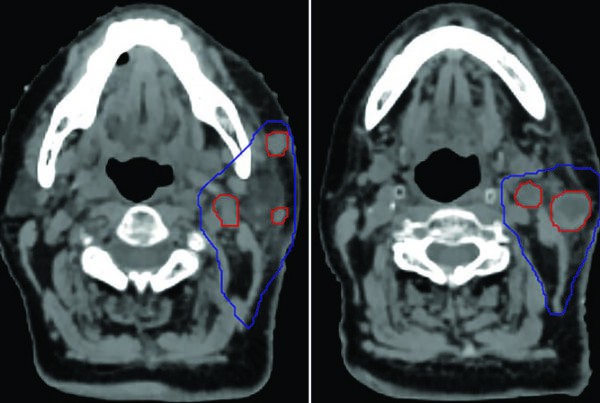
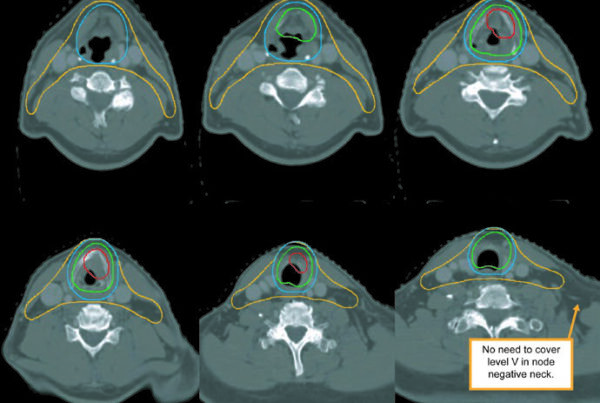
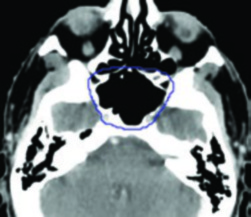
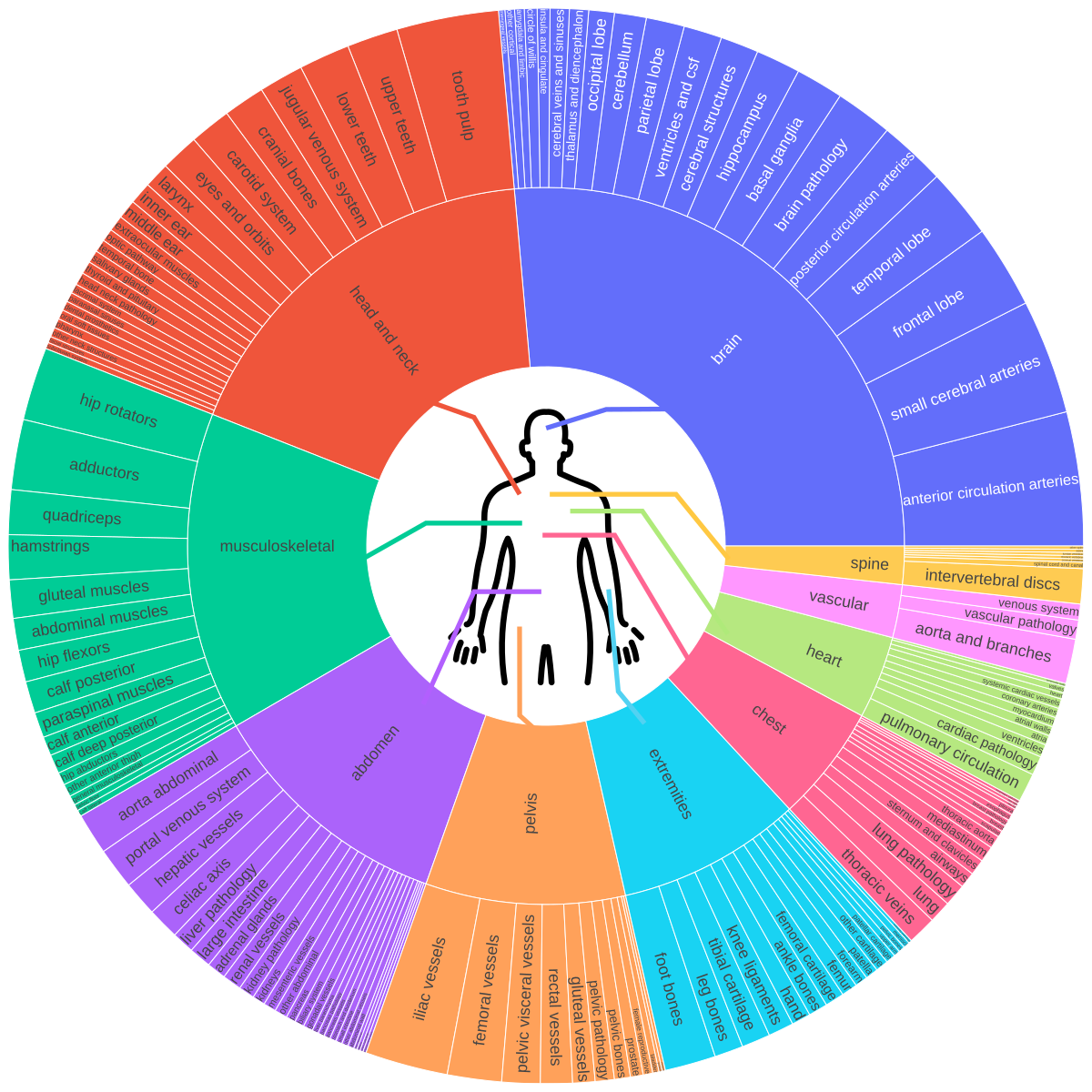
The original paper (arXiv:2511.11450) documents training on 158 public datasets covering brain, head & neck, thorax, abdomen, pelvis, and musculoskeletal system — including vascular structures, organ substructures, and lesions. A foundation that reflects AI’s migration from isolated algorithms to workflow integration.
Web Interface: 3D Viewer, RTStruct, and Low-VRAM Engineering
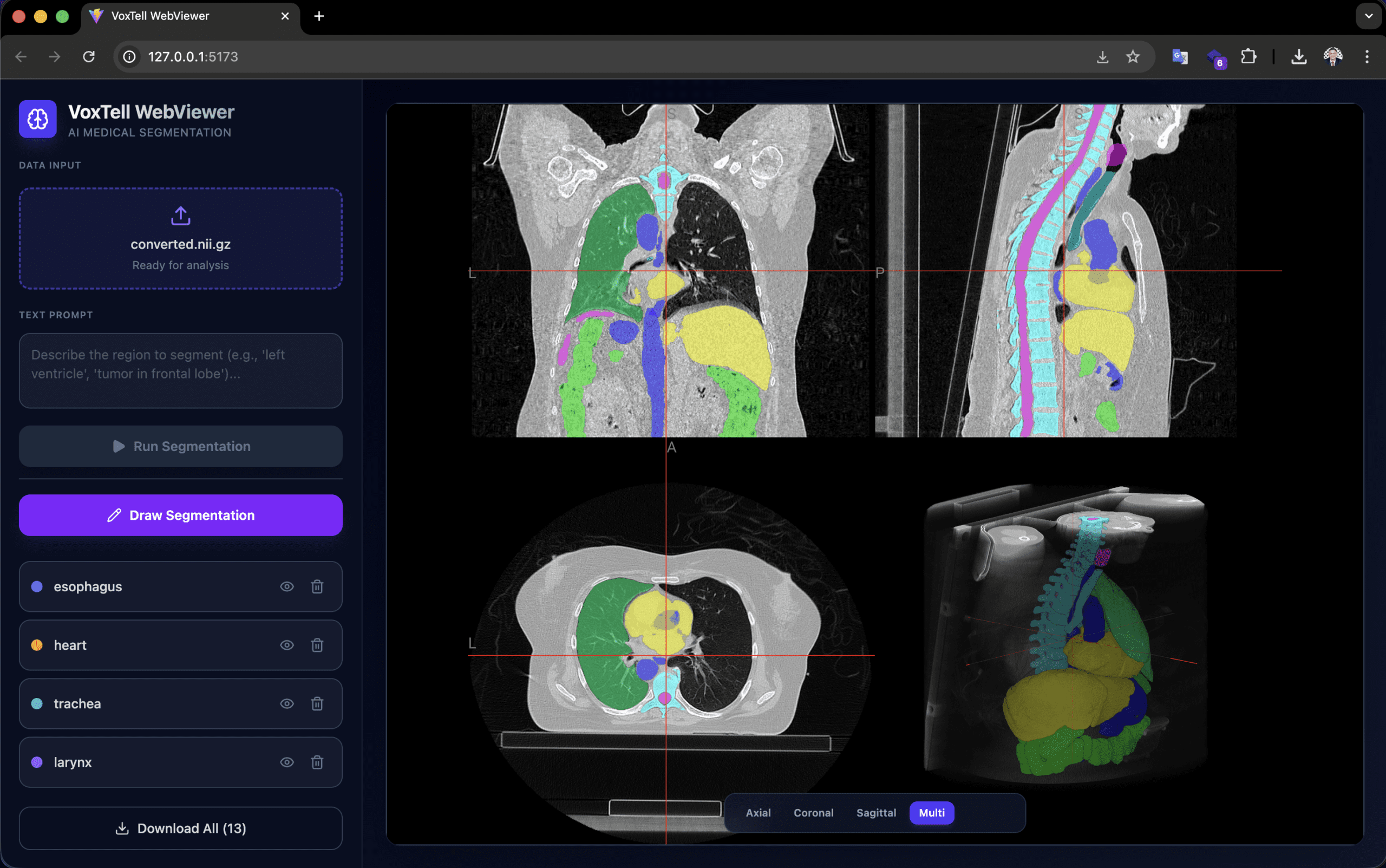
The voxtell-web-plugin is a FastAPI + React/TypeScript application that puts the model behind an accessible interface. The operator uploads a volume (.nii, .nii.gz, or DICOM), types a prompt like “liver” or “prostate tumor,” and receives the 3D mask overlaid on the NiiVue viewer in real time.
Low-VRAM engineering is the practical differentiator. The Qwen3-Embedding-4B text encoder runs in float16, reducing memory usage from ~15 GB to ~7.5 GB. The memory allocator uses expandable_segments=True to reduce fragmentation, and the sliding window operates with perform_everything_on_device=False for partial CPU offload. This means 12 GB GPUs can already run inference — hardware found in research workstations, not just clusters.
The viewer supports accumulation of multiple segmentations (liver + spleen + kidneys in the same session), manual drawing for refinement, and export in NIfTI and RTStruct. RTStruct export is particularly relevant: it produces a DICOM-RT file that can be imported into treatment planning systems for comparative evaluation — always in a research context.
Orientation warning: images must be in RAS orientation for correct left/right anatomical localization. Orientation mismatches produce mirrored or incorrect results. PyTorch 2.9.0 has an OOM bug in 3D convolutions; version 2.8.0 or earlier is recommended.
Varian Eclipse ESAPI: How the Integration Works
The VoxTell-ESAPI adds two components to the ecosystem: a Python/FastAPI server that receives CT data over HTTP and runs GPU inference, and a C# ESAPI plugin that extracts CT from Varian Eclipse, sends it to the server, and reimports resulting contours as RT structures.
The complete workflow:
- Operator opens a patient in Eclipse with CT and existing structure set
- Plugin creates a session on the server, sending volume geometry (origin, row/column/slice direction, spacing)
- For each Z slice, voxels are extracted as
ushort[xSize, ySize], widened to int32, serialized in little-endian, gzip-compressed, and base64-encoded — reducing payload ~4× - After all slices are sent, the server assembles the NIfTI volume with LPS→RAS conversion
- Operator types prompts (e.g., “liver, left kidney, spleen”) and submits
- Inference runs asynchronously — Eclipse UI is not blocked
- Server extracts 2D contours from masks and returns coordinates in LPS (patient)
- Plugin imports via
structure.AddContourOnImagePlane(contour_points_lps, z_index)
Existing structures are matched by name (exact, case-insensitive, or fuzzy). Missing structures are auto-created with DICOM type CONTROL. Names are sanitized to 16 characters (e.g., “left kidney” → “left_kidney”).
This plugin is intended exclusively for non-clinical environments: ECNC (External Calculation and Non-Clinical) and Varian TBOX (training box). It must never be used in a clinical environment.
DICOM Coordinate Conversion Pipeline: LPS, RAS, and the Math
Coordinate conversion between DICOM (LPS) and NIfTI (RAS) is the most critical technical point of the entire integration. An error at this stage produces mirrored volumes, anteroposterior-inverted contours, or structures on the wrong side of the patient. The pipeline implements the transformation rigorously.
DICOM Geometry → LPS Affine
Eclipse exposes image geometry (origin, row direction, column direction, slice direction, spacing). The server constructs the 4×4 affine matrix mapping voxel indices to millimetre positions in DICOM LPS (Left, Posterior, Superior):
$$x_{LPS} = A_{LPS} \begin{bmatrix} i \\ j \\ k \\ 1 \end{bmatrix}$$
Where the columns of $A_{LPS}$ are:
- Column 0:
row_direction × x_res(column +X axis) - Column 1:
col_direction × y_res(row +Y axis) - Column 2:
slice_direction × z_res(slice +Z axis) - Column 3:
origin(position of voxel 0,0,0)
LPS → RAS Conversion
DICOM and NIfTI use opposite conventions on the first two axes:
| System | X | Y | Z |
|---|---|---|---|
| DICOM/Eclipse (LPS) | Patient Left | Patient Posterior | Patient Superior |
| NIfTI/VoxTell (RAS) | Patient Right | Patient Anterior | Patient Superior |
The transformation requires inverting the first two axes:
$$A_{RAS} = \operatorname{diag}(-1,-1,1,1) \cdot A_{LPS}$$
In code, the volume is transposed from (Z,Y,X) to (X,Y,Z) for NIfTI convention, and the X and Y affine axes are inverted. A naive copy produces a mirrored, anteroposterior-inverted volume — exactly the kind of error that surfaces only in rigorous clinical review, not in automated tests.
Return Path: RAS Masks → LPS Contour Points
After inference, the inverse path uses scikit-image’s find_contours to extract 2D contour lines per slice, projecting voxel indices back to LPS millimetres using the session’s stored affine:
$$\text{pts}_{LPS} = (\text{vox\_coords} \cdot A_{LPS}^T)[:, :3]$$
Points are sent to Eclipse, which applies them directly via AddContourOnImagePlane().
Evaluation Metrics
Two standard metrics evaluate segmentation quality:
The Dice coefficient measures overlap between predicted segmentation $X$ and reference $Y$:
$$DSC(X,Y) = \frac{2|X \cap Y|}{|X| + |Y|}$$
Hausdorff distance measures the worst-case point-to-point divergence between surfaces:
$$HD(X,Y) = \max\left\{\sup_{x \in X}\inf_{y \in Y} d(x,y),\; \sup_{y \in Y}\inf_{x \in X} d(x,y)\right\}$$
Research Plugins, SaMD Boundaries, and Why Regulatory Language Matters

The medical software market operates under rigorous regulation. Any software that influences diagnostic or therapeutic decisions may be classified as Software as a Medical Device (SaMD), subject to frameworks like IEC 62304, ISO 14971, IMDRF, and regulations from agencies such as FDA, ANVISA, and CE Marking.
The plugins described in this article — web and ESAPI — are research, experimentation, prototyping, and technical evaluation tools. Specifically:
- The original VoxTell model is work by the research group cited in the paper (Rokuss et al., 2025), not by RT Medical Systems
- RT Medical contributes open source engineering of public integrations and extensions around VoxTell
- The ESAPI plugin is intended exclusively for ECNC and Varian TBOX — non-clinical environments
- These plugins must never be used clinically
- They are not approved, cleared, validated, or authorized medical software by any regulatory agency
- There is no formal endorsement from Varian, DKFZ, MIC-DKFZ, or the original paper authors
Clinical use of any AI-assisted segmentation tool would require independent validation, a quality management system, risk analysis (ISO 14971), cybersecurity review, and full regulatory assessment. These are not formalities — they are the barriers separating research prototypes from devices that influence patient treatment.
For professionals working with DICOM software development or DICOM infrastructure implementation, understanding this boundary is essential before evaluating any AI tool.
Integration Engineering: What Radiotherapy Demands from Software
The technical value of these integrations is not in the model itself — segmentation models appear every quarter. The value lies in demonstrating the engineering competencies that any radiotherapy software company must master:
- DICOM interoperability: bidirectional format conversion (NIfTI ↔ DICOM), affine manipulation, volume orientation, RTStruct export
- TPS integration: ESAPI communication, voxel serialization, contour import in patient coordinates
- Resource optimization: consumer GPU inference, CPU offload, payload compression
- Asynchronous workflow: TTL sessions, non-blocking polling, cancellation and cleanup
- Governance: clear separation between research and clinical product, precise regulatory language
Each of these is a real requirement in projects like RTConnect and contour review pipelines — not theoretical exercises, but problems that arise in every integration with actual equipment and planning systems. TG-263 structure standardization is another direct point of convergence.
Next Steps and Context for Teams
VoxTell’s public roadmap indicates fine-tuning support has not yet been released. When available, it will open possibilities for adapting to specific structures of interest — for example, head and neck OAR structures per institutional protocols — again in a research context.
If your team is evaluating AI-assisted contouring workflows, validation pipelines, or review and governance layers around segmentation, RT Medical Systems can help structure that discussion.
All technical information in this article was extracted from public sources: the VoxTell paper (arXiv:2511.11450, Rokuss et al., 2025) and the GitHub repositories gomesgustavoo/voxtell-web-plugin and gomesgustavoo/VoxTell-ESAPI.